Data Visualization Galleries
Publish date: Nov 13, 2021Palletes
Show all palettes. - Diverging Scale (BrBG, PiYG, PRGn, PuOr, RdBu, RdGy, RdYlBu, RdYlGn, Spectral) - Qualitative Scale (Accent, Dark2, Paired, Pastel1, Pastel2, Set1, Set2, Set3) - Sequential Scale (Blues, BuGn, BuPu, GnBu, Greens, Greys, Oranges, OrRd, PuBu, PuBuGn, PuRd, Purples, RdPu, Reds, YlGn, YlGnBu, YlOrBr, YlOrRd)
library(RColorBrewer)
display.brewer.all() # colorblindFriendly = TRUE will exclude some palettes.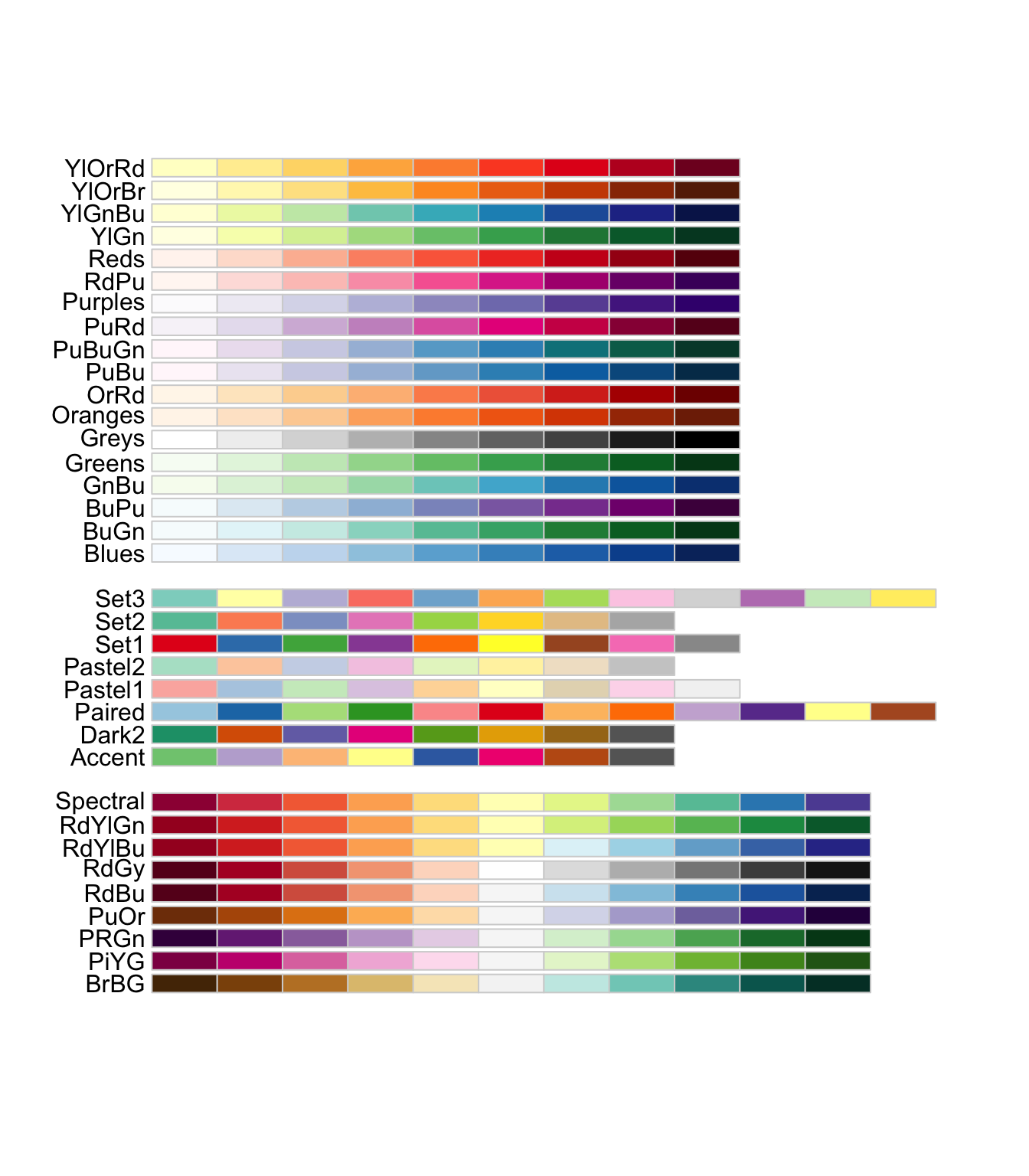
# Use following functions to apply preset or our own palette
# scale_fill_brewer(palette = "Set2")
# scale_color_brewer(palette = "Set2")
# scale_fill_manual(values=c('#00429d', '#4771b2', '#73a2c6', '#a5d5d8', '#ffffe0'))Fonts
library(extrafont)
library(tidyverse)
library(patchwork)
# font_import()
plot <- ggplot(mtcars, aes(x=wt, y=mpg)) +
geom_point() +
labs(
title = "Fuel Efficiency of 32 Cars",
x= "Weight (x1000 lb)",
y = "Miles per Gallon"
)+
theme_minimal()+
NULL
p1 <- plot + theme(text=element_text(size=16, family="Montserrat"))
p2 <- plot + theme(text=element_text(size=16, family="Bahnschrift"))
p3 <- plot + theme(text=element_text(size=16, family="Oswald"))
p4 <- plot + theme(text=element_text(size=16, family="Rock Salt"))
# Use patchwork to arrange 4 graphs on same page
(p1 + p2) / (p3 + p4)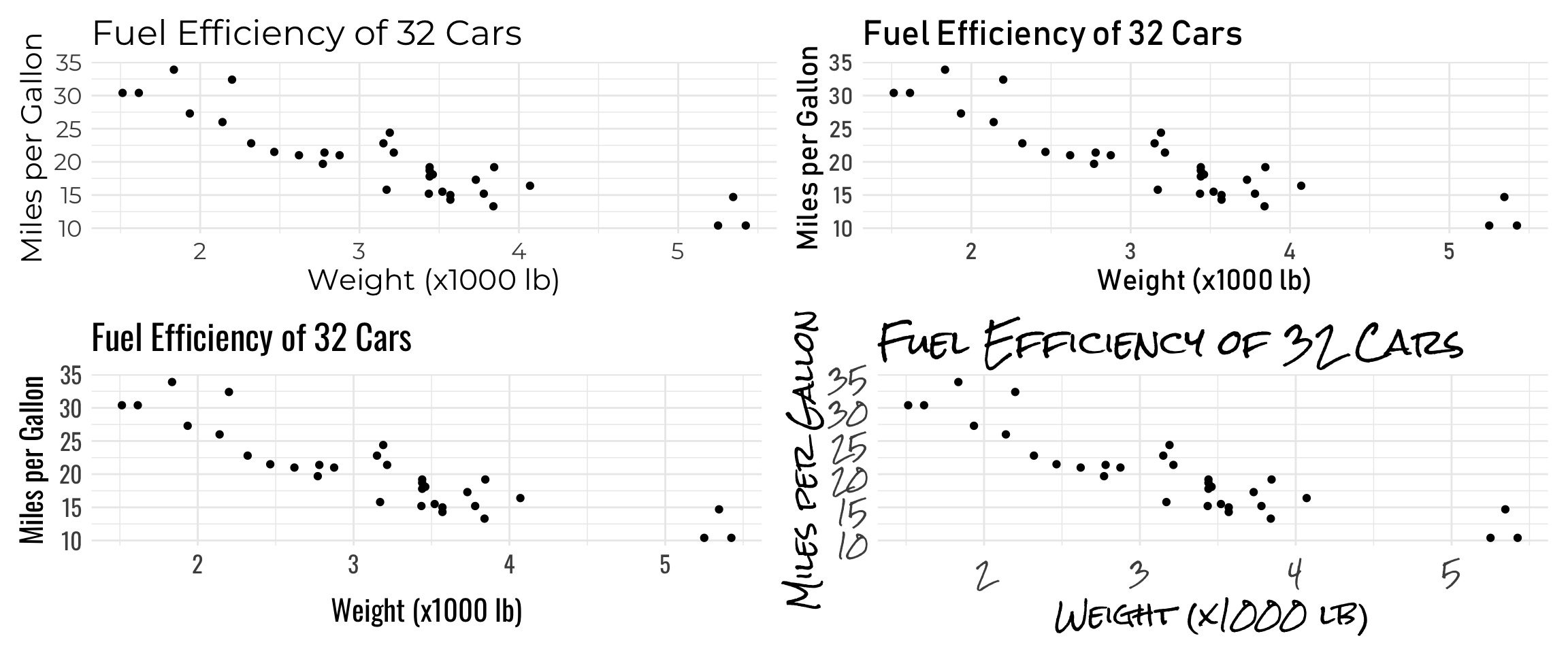
Bar plot - Emperors
library(tidyverse)
library(vroom)
library(extrafont)
library(ggtext)# emperors data
url <- "https://github.com/rfordatascience/tidytuesday/raw/master/data/2019/2019-08-13/emperors.csv"
emperors <- vroom(url)
# Count of Causes of Death
# New variable 'assassination' to colour differently (Highlight)
emperors_assassinated <- emperors %>%
count(cause) %>%
arrange(n) %>%
mutate(
assassination = ifelse(cause == "Assassination" | cause == "Execution", TRUE, FALSE),
cause = fct_inorder(cause)
)Trick: use fct_inorder after arrange to plot in a proper order.
# get hex code for color
gplots::col2hex("grey80")## [1] "#CCCCCC"gplots::col2hex("tomato")## [1] "#FF6347"Trick: gplots::col2hex to get hex code for color
# define colours to use: grey for everything, tomato for assassination
my_colours <- c("#CCCCCC", "#FF6347")
# let us create a text label to add as annotation to our graph
label <- "Execution is a third as likely \n as assassination"
emperors_assassinated %>%
ggplot(aes(x = n, y = cause, fill = assassination)) +
geom_col() +
scale_fill_manual(values = my_colours)+
geom_text(
aes(label = n, x = n - .25),
colour = "white",
size = 5,
hjust = 1,
family="Lato"
) +
# change font colour
labs(title = "<b> Cause of death of Roman emperors</b><br>
<span style = 'font-size:12pt'>Roughly half of the emperors died of <span style='color:#FF6347'>assassination</span> and <span style='color:#FF6347'>execution </span>.</span>",
x= "Number of emperors",
y = "")+
#add an arrow to draw attention to a value
geom_curve(
data = data.frame(x = 15, y = 3.2, xend = 8.1, yend = 5),
mapping = aes(x = x, y = y, xend = xend, yend = yend),
colour = "grey15",
size = 0.5,
curvature = 0.25,
arrow = arrow(length = unit(2, "mm"), type = "closed"),
inherit.aes = FALSE
) +
# add the text label on the graph
geom_text(
data = data.frame(x = 15, y = 3, label = label),
aes(x = x, y = y, label = label),
colour="#FF6347",
family="Lato",
hjust = 0.5,
lineheight = .8,
inherit.aes = FALSE,
) +
theme_minimal() +
theme(
plot.title.position = "plot",
plot.title = element_textbox_simple(size=16), # with this line, html is compiled
axis.text = element_text(size=12),
legend.position = "none") +
NULL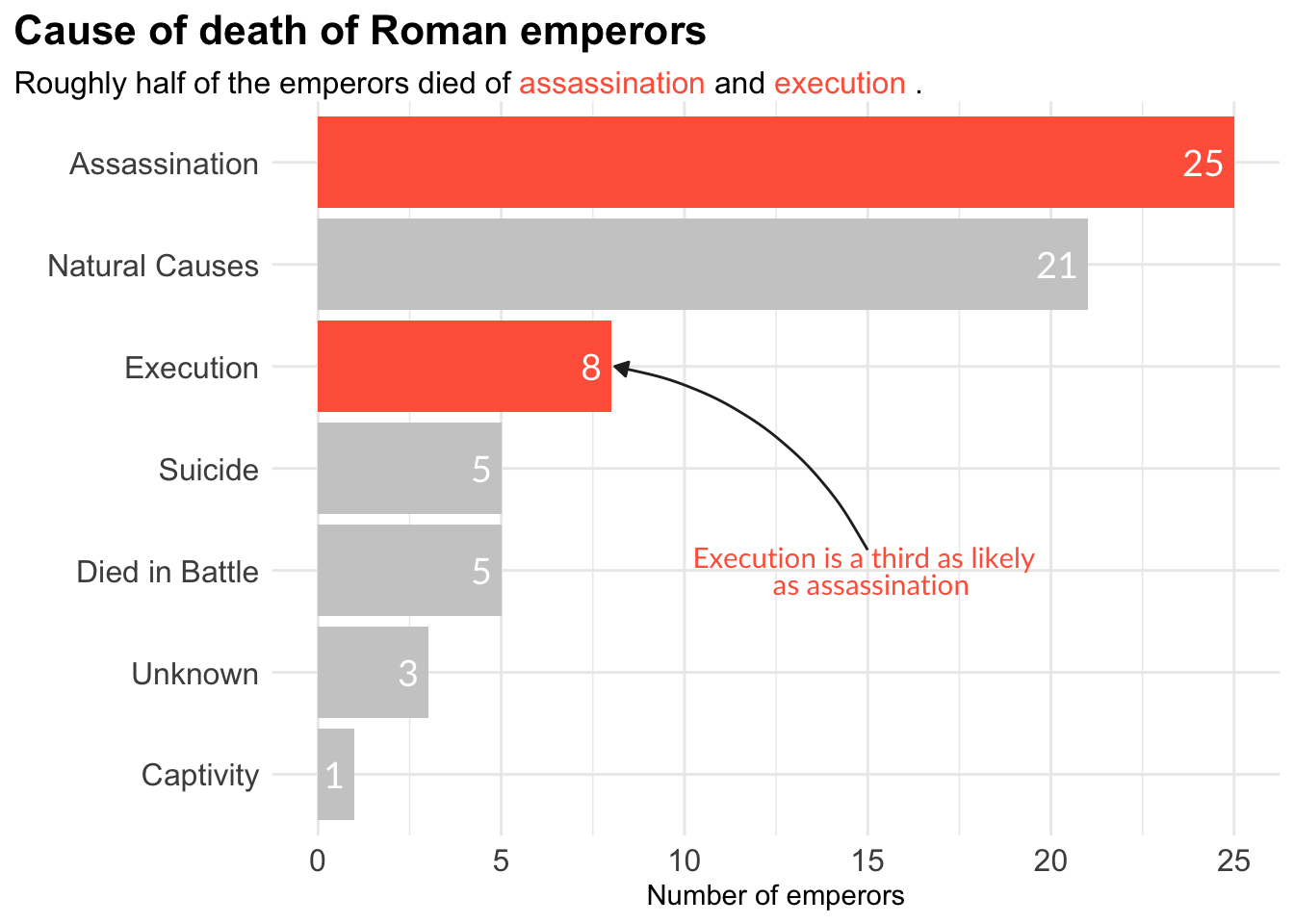
Treemap - Car Manufacturers
library(tidyverse)
library(treemapify)
library(ggalluvial)
library(ggridges)
library(ggthemes)# Treemaps
# further treemap examples https://github.com/wilkox/treemapify
mpg %>%
filter(year == 1999) %>%
count(manufacturer) %>%
ggplot(aes(area = n,
fill = manufacturer,
label = manufacturer)) +
geom_treemap() +
geom_treemap_text() +
theme(legend.position = "none")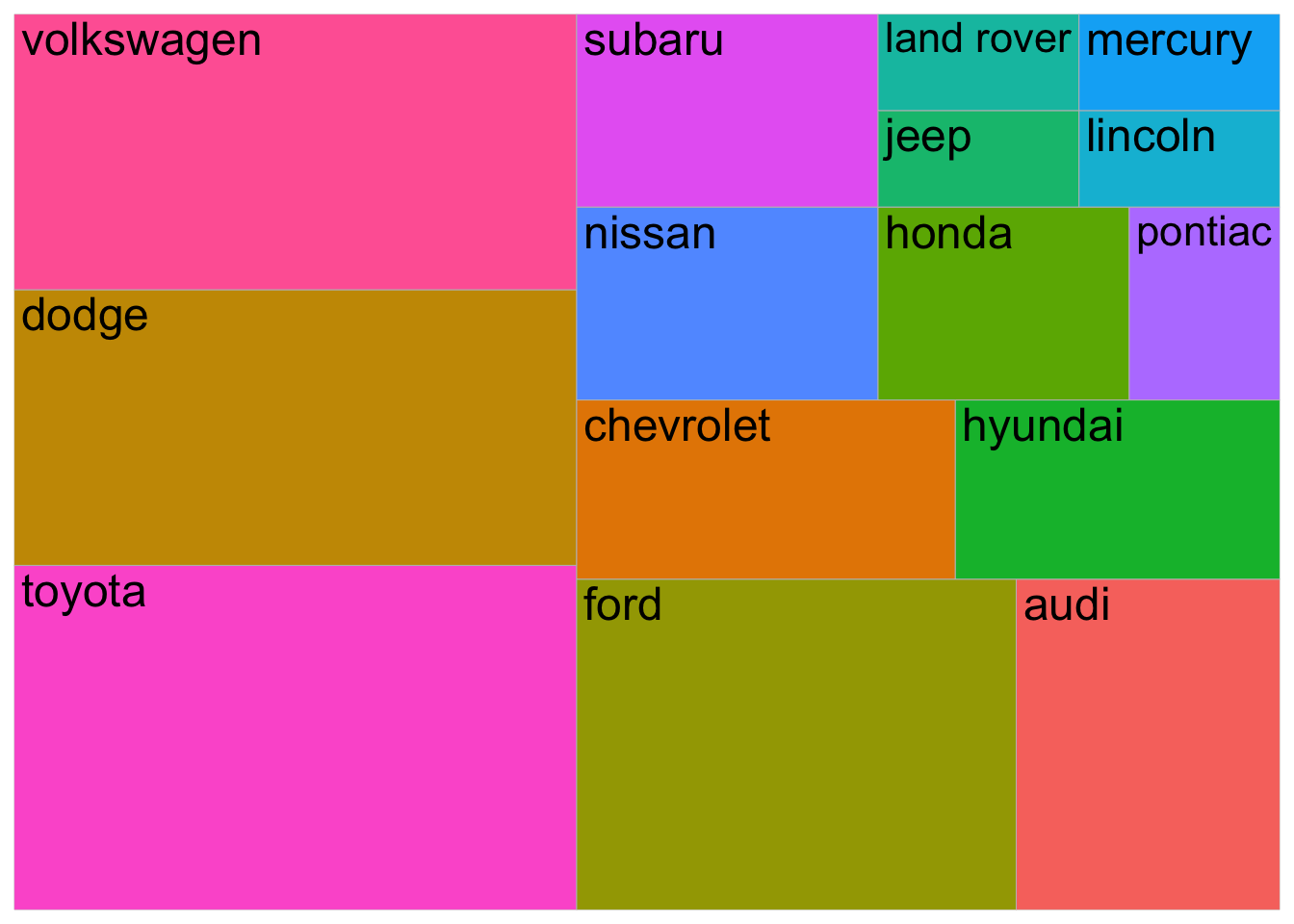
Alluvial Plot - Vaccination Survey
# Alluvial or flow/Sankey diagrams
# https://corybrunson.github.io/ggalluvial/articles/ggalluvial.html
data(vaccinations)
ggplot(vaccinations,
aes(x = survey, y = freq,
alluvium = subject, stratum = response,
fill = response, label = response)) +
scale_x_discrete(expand = c(.1, .1)) +
geom_flow() +
geom_stratum(alpha = .5) +
geom_text(stat = "stratum", size = 3) +
theme(legend.position = "none") +
labs(title = "Vaccination survey response at three times")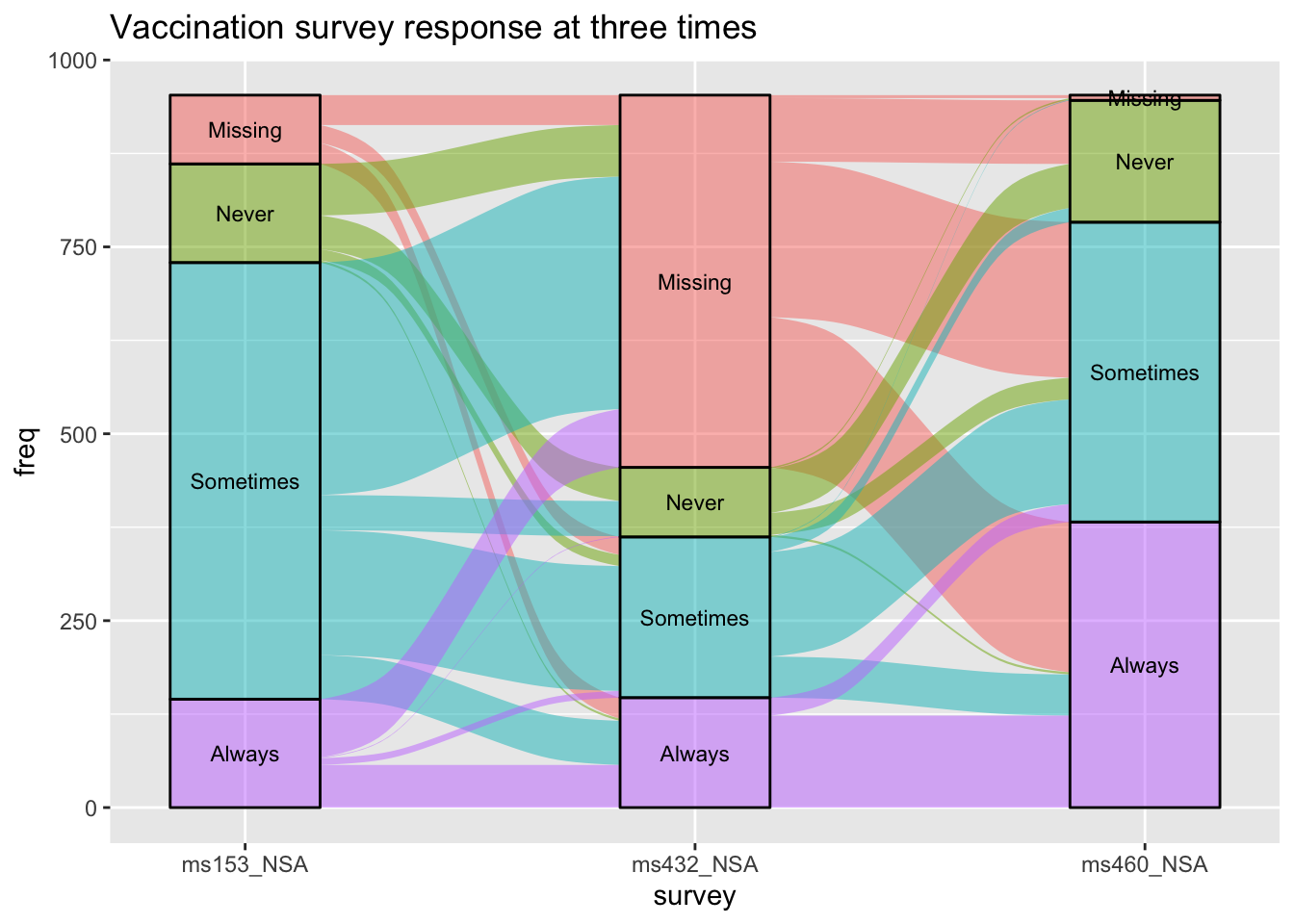
Density Plot
1-Dimension: Distributions
library(gapminder)
# create gapminder2007, by just looking at 2007 data
gapminder2007 <- filter(gapminder,
year == 2007)
# use ggridges::geom_density_ridges() for multiple density plots
ggplot(filter(gapminder2007,
continent != "Oceania"),
aes(x = lifeExp,
fill = continent,
y = continent)) +
geom_density_ridges(alpha = 5/8)+
theme_minimal()+
guides(fill=FALSE)+
NULL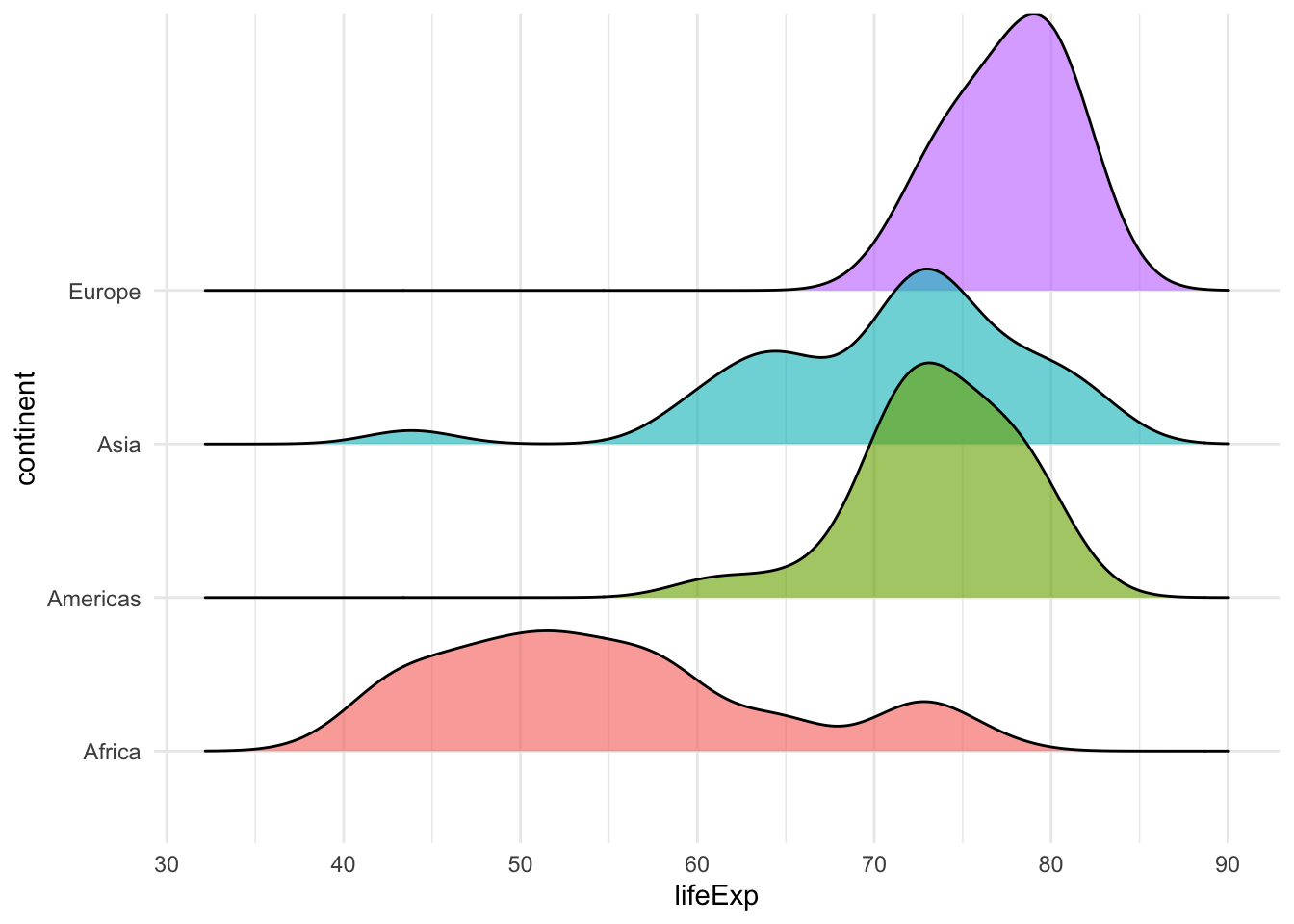
Comparision - With background
library(vroom)
# load titanic data
titanic <- vroom(here::here("data", "titanic3.csv"))
# Define a vector of colours... grey80 for background, and two different colours for male/female
density_colours <- c(
"male" = "#0072B2",
"female" = "#D55E00",
"all passengers" = "grey80"
)
titanic %>%
drop_na(sex, age) %>% # remove NAs for sex and age, and pass dataframe to ggplot
# in the ggplot(), besides `x = age`, add `y = ..count..` to `aes()`
ggplot(aes(x = age, y = ..count..)) +
# Add an additional `geom_density()` that will draw the background density, for all participants.
# This should come *before* the 'normal' `geom_density()` so that it draws the background.
geom_density(
# In the background geom_density(), set the `data` argument to be a function. This function
# takes a data frame and removes sex (select x, -sex) (on which sex we will facet on later).
data = function(x) select(x, -sex),
# Set both `colour` and `fill` equal to "all passengers", *not* `sex`.
aes(fill = "all passengers", colour = "all passengers")
) +
# now that we have the grey background, we can plot densities
# coloured and facet_wrapped by sex
geom_density(aes(fill = sex, colour = sex), alpha = 0.7, bw = 2) +
# `facet_wrap()` to facet the plot by `sex`.
# labeller() makes it easy to assign different labellers to different factors
facet_wrap(~sex, labeller = labeller(sex = function(sex) paste(sex, "passengers"))) +
# use the colours you defined for male, female, and all participants
scale_colour_manual(
values = density_colours,
breaks = c("male", "female", "all passengers"),
labels = c( "males", "females","all passengers"),
name = NULL,
guide = guide_legend(direction = "horizontal")
)+
scale_fill_manual(
values = density_colours,
breaks = c("male", "female", "all passengers"),
labels = c( "males", "females","all passengers"),
name = NULL,
guide = guide_legend(direction = "horizontal")
)+
scale_x_continuous(name = "age (years)", limits = c(0, 75), expand = c(0, 0)) +
scale_y_continuous(limits = c(0, 40), expand = c(0, 0), name = "scaled density") +
coord_cartesian(clip = "off") +
theme_minimal() +
theme(legend.position = "bottom",
legend.justification = "center",
text=element_text(size=12, family="Lato"),
plot.title.position = "plot"
) +
NULL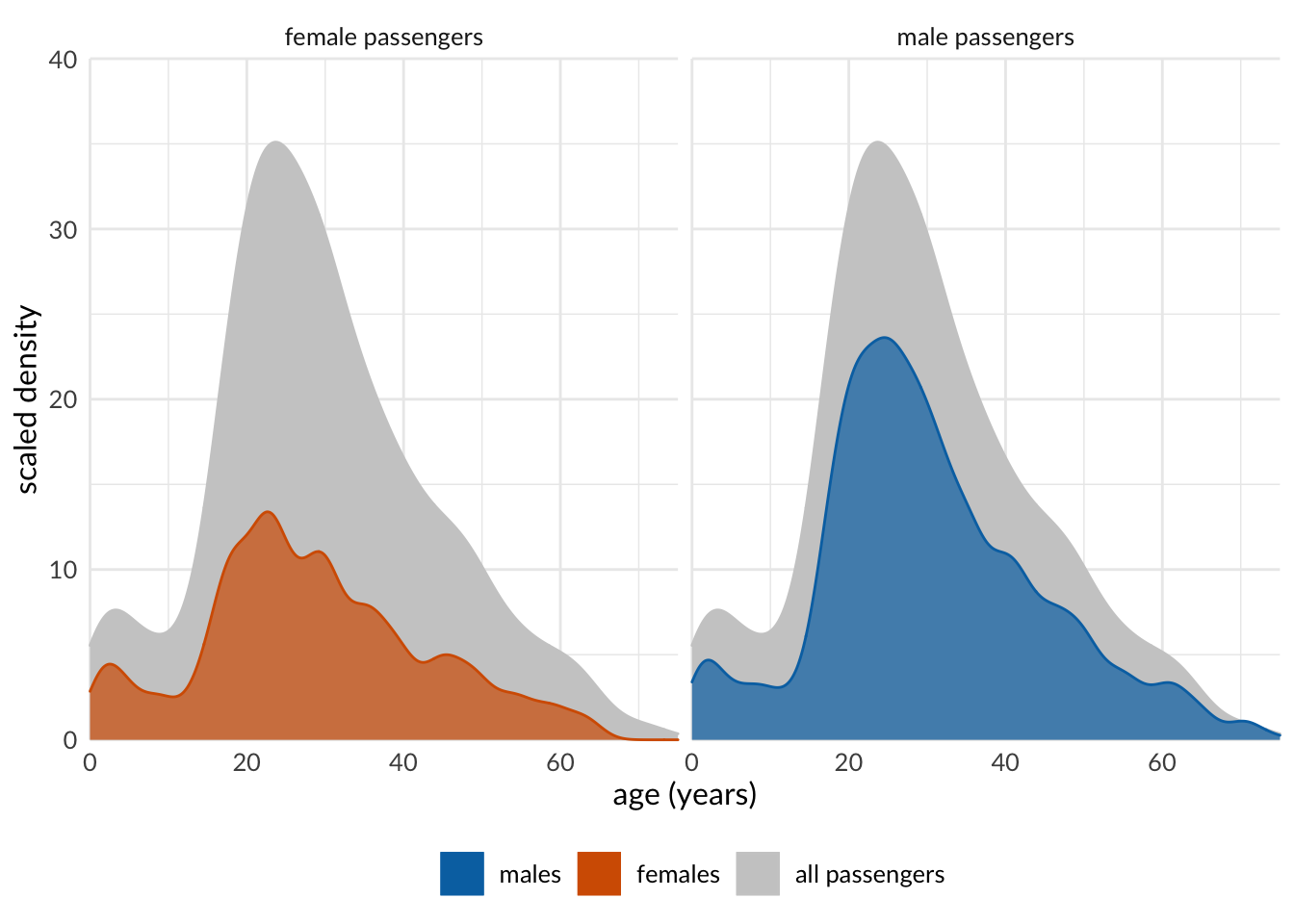
Comparision - with half box plot
library(tidyverse)
library(gapminder)
library(ggridges)
library(gghalves)
# use gghalves::geom_half_boxplot(), gghalves::geom_half_point()
ggplot(filter(gapminder2007,
continent != "Oceania"),
aes(y = lifeExp,
x = continent,
colour = continent)) +
geom_half_boxplot(side = "l") + # half boxplot to the left
geom_half_point(side = "r")+ # points to the right
theme_minimal()+
guides(fill = FALSE, color = FALSE)+
NULL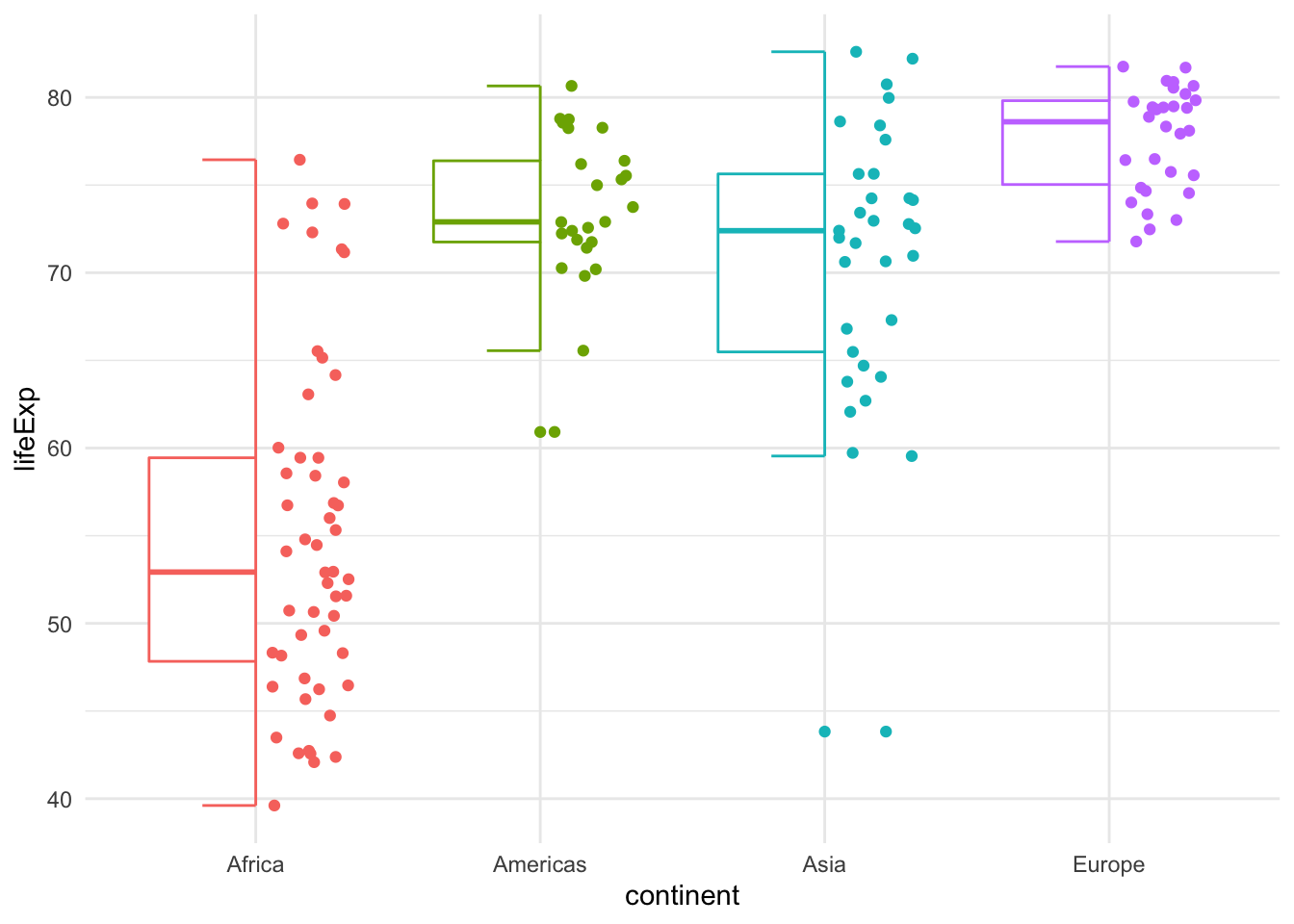
# Raincloud plots
ggplot(filter(gapminder2007,
continent != "Oceania"),
aes(y = lifeExp,
x = continent,
colour = continent)) +
geom_half_point(side = "l", size = 0.3) +
geom_half_boxplot(side = "l", width = 0.5,
alpha = 0.3, nudge = 0.1) +
geom_half_violin(aes(fill = continent),
alpha = 0.8,
side = "r") +
guides(fill = FALSE, color = FALSE) +
coord_flip()+
theme_minimal()+
NULL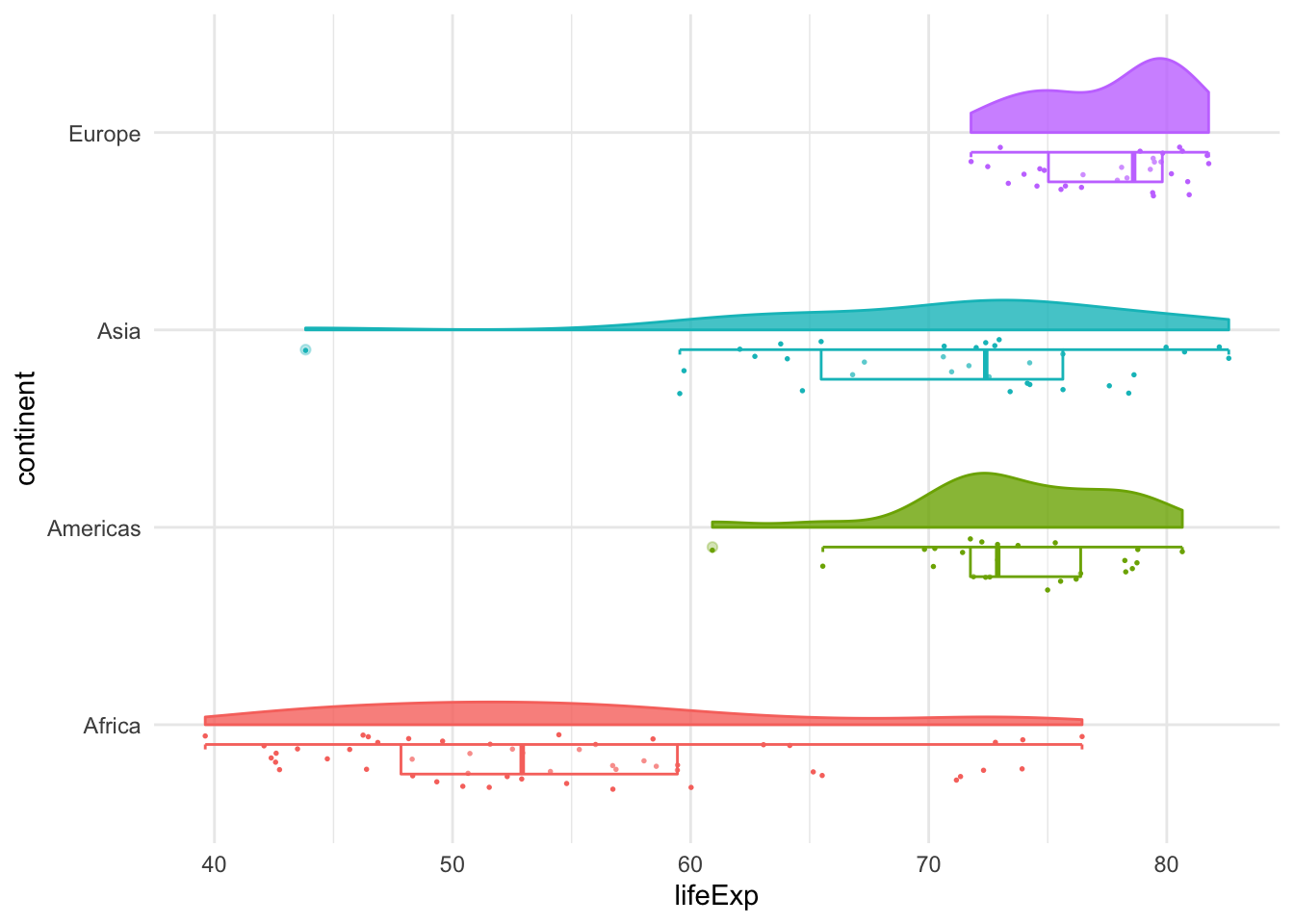
2-Dimension
mpg %>%
filter(class != "2seater") %>%
ggplot(aes(x = cty, y = hwy)) +
geom_density_2d(aes(color = class)) +
facet_wrap(~class) +
theme(legend.position = "none")+
labs(x="city miles per gallon",
y="highway miles per gallon")+
NULL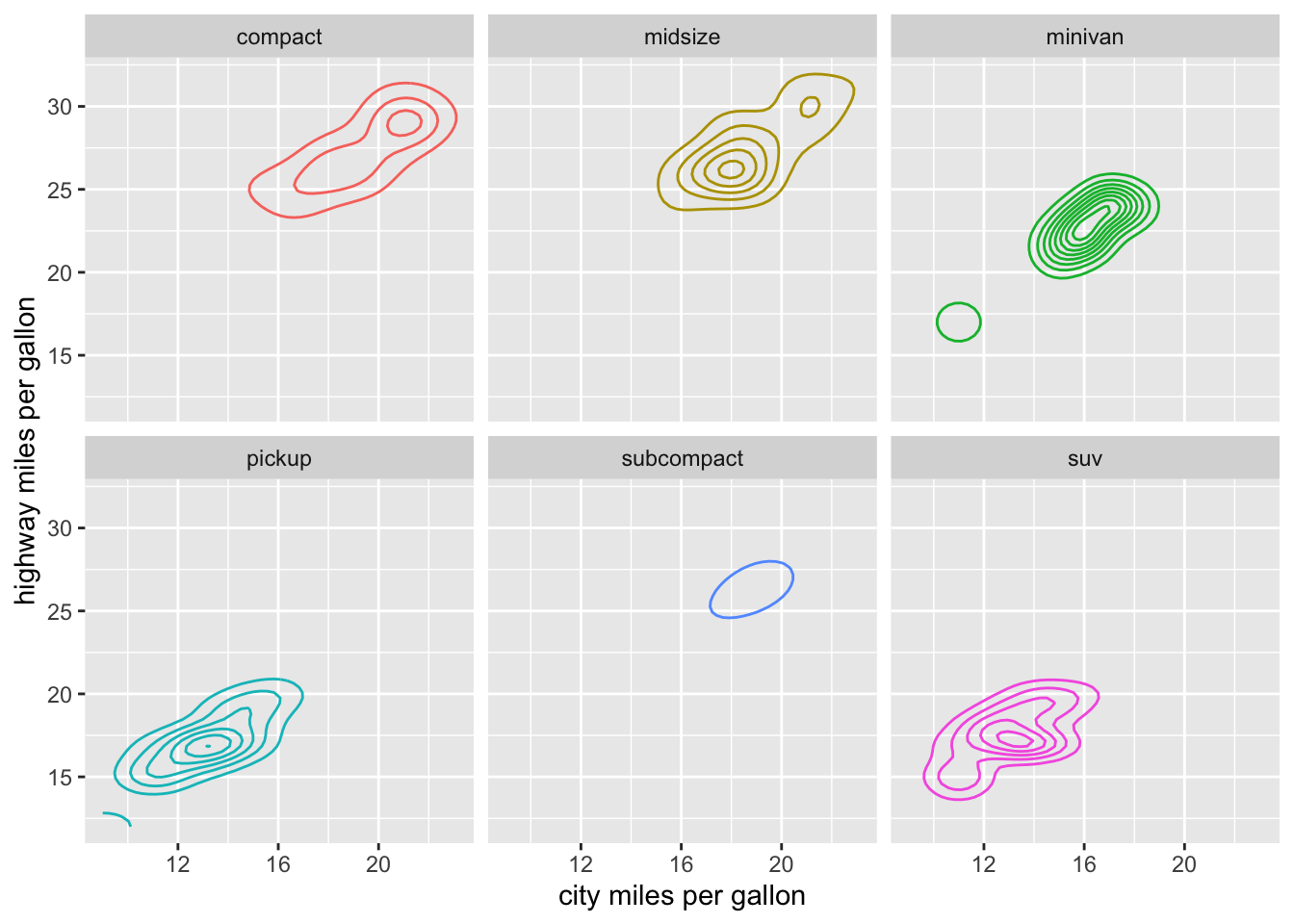
# geom_hex: Divides the plane into regular hexagons
mpg %>%
filter(class != "2seater") %>%
ggplot(aes(x = cty, y = hwy)) +
geom_hex(aes(color = class), bins = 10) +
facet_wrap(~class) +
#theme(legend.position = "none")+
labs(x="city miles per gallon",
y="highway miles per gallon")+
NULL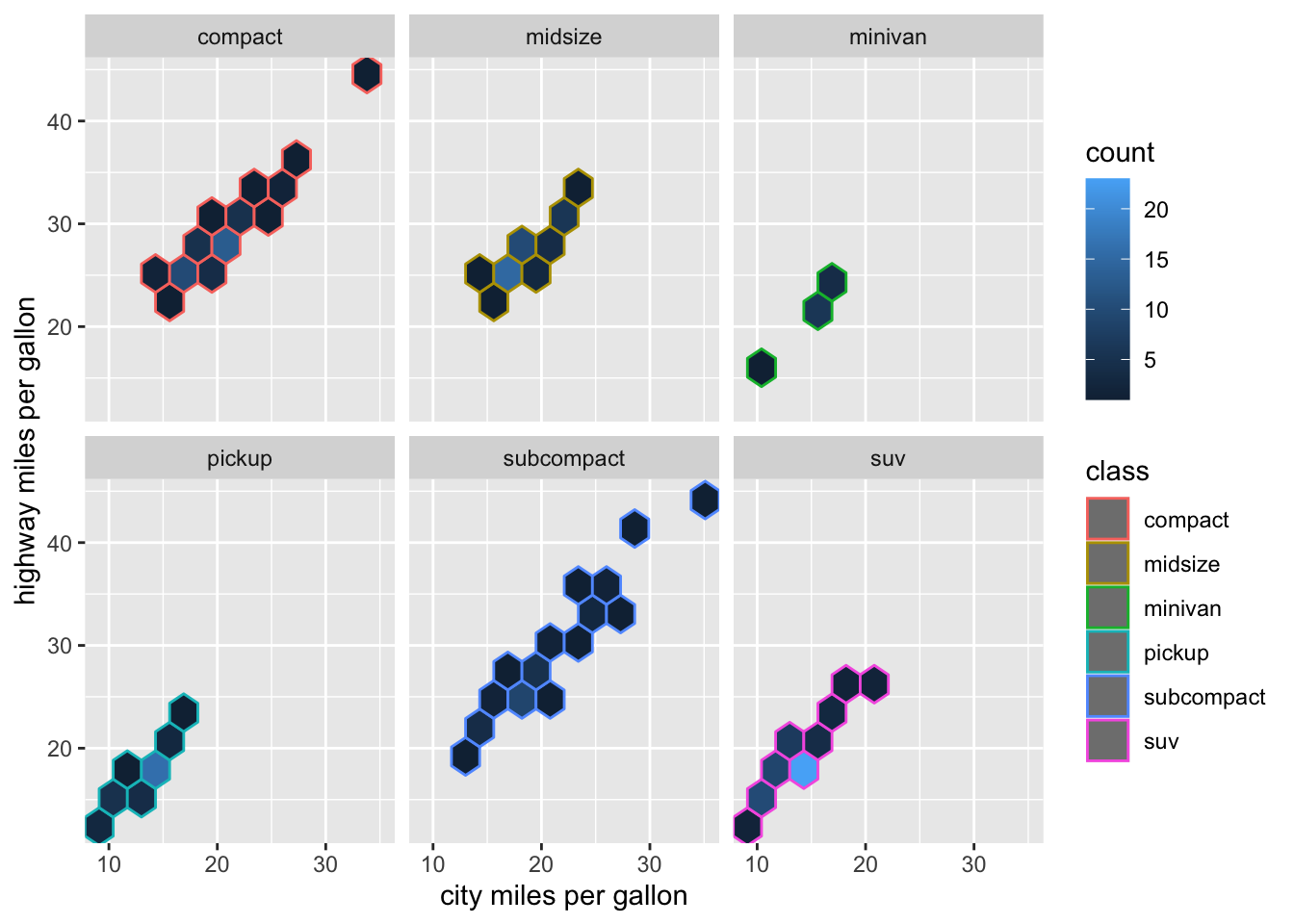
Heatmap
library(tidyverse)
library(wbstats)
# https://data.worldbank.org/indicator/IT.NET.USER.ZS
# Download data for Individuals Using Internet (% Of Population) IT.NET.USER.ZS
# using the wbstats package
internet <- wb_data(country = "countries_only",
indicator = "IT.NET.USER.ZS",
start_date = 1995, end_date = 2017)
country_list = c("United States", "China", "India", "Japan", "Algeria",
"Brazil", "Germany", "France", "United Kingdom", "Italy", "New Zealand",
"Canada", "Mexico", "Chile", "Argentina", "Norway", "South Africa", "Kenya",
"Israel", "Iceland")
internet_short <- filter(internet, country %in% country_list) %>%
rename(value = IT.NET.USER.ZS) %>%
mutate(users = ifelse(is.na(value), 0, value),
year = as.numeric(date))
internet_summary <- internet_short %>%
group_by(country) %>%
summarize(year1 = min(year[users > 0]),
last = users[n()]) %>%
arrange(last, desc(year1))
internet_short <- internet_short %>%
mutate(country = factor(country, levels = internet_summary$country))
ggplot(filter(internet_short, year > 1993),
aes(x = year, y = country, fill = users)) +
geom_tile(color = "white", size = 0.25) +
scale_fill_viridis_c(
option = "A", begin = 0.05, end = 0.98,
limits = c(0, 100),
name = "internet users / 100 people",
guide = guide_colorbar(
direction = "horizontal",
label.position = "bottom",
title.position = "top",
ticks = FALSE,
barwidth = grid::unit(3.5, "in"),
barheight = grid::unit(0.2, "in")
)
) +
scale_x_continuous(expand = c(0, 0), name = NULL) +
scale_y_discrete(name = NULL, position = "right") +
theme(
axis.line = element_blank(),
axis.ticks = element_blank(),
axis.ticks.length = grid::unit(1, "pt"),
legend.position = "top",
legend.justification = "left",
legend.title.align = 0.5,
legend.title = element_text(size = 12*12/14)
)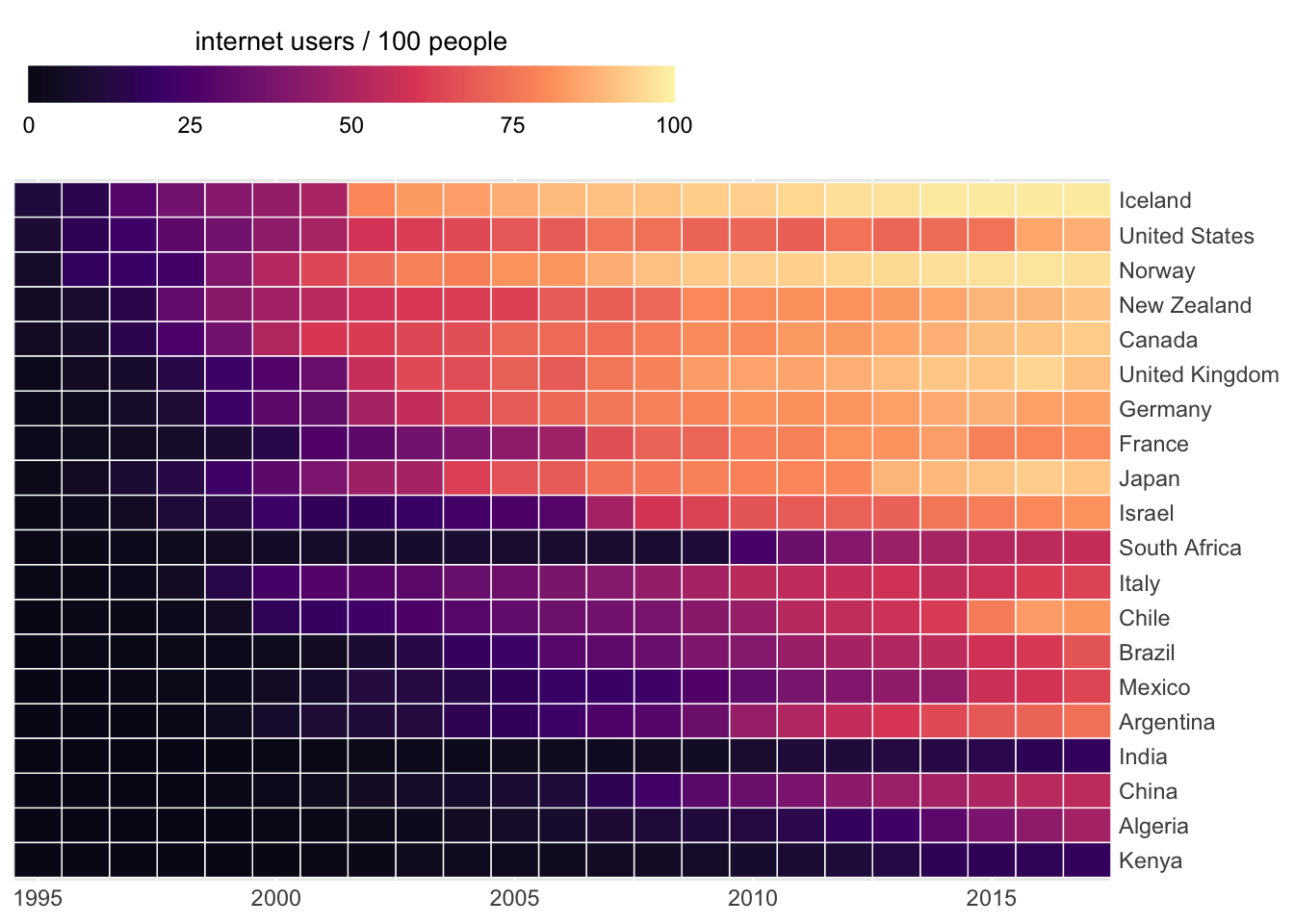 ## X-Y relationships
## X-Y relationships
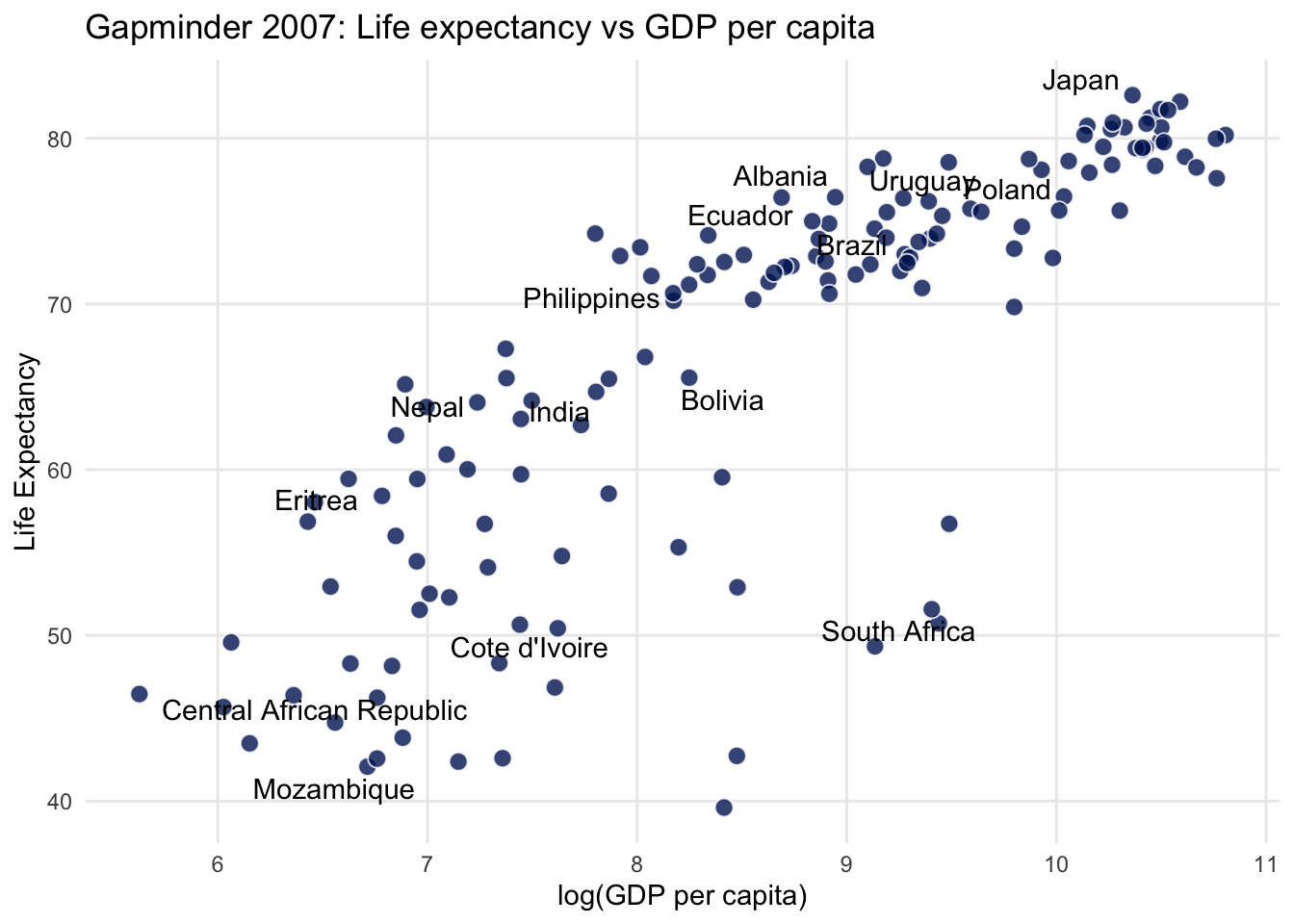
Linear Relationship
City miles vs Highway miles per gallon
library(tidyverse)
library(gapminder)
library(extrafont)
library(ggrepel)set.seed(7)
# 20 random countries
some_countries <- gapminder$country %>%
levels() %>%
sample(20)
gapminder %>%
filter(year == 2007) %>%
mutate(
label = ifelse(country %in% some_countries, as.character(country), "")
) %>%
ggplot(aes(log(gdpPercap), lifeExp)) +
geom_point(
size = 3,
alpha = 0.8,
shape = 21,
colour = "white",
fill = "#001e62"
)+
geom_text_repel(aes(label=label))+
theme_minimal()+
theme(panel.grid.minor = element_blank())+
labs(
title = "Gapminder 2007: Life expectancy vs GDP per capita",
x = "log(GDP per capita)",
y = "Life Expectancy"
)
Secondary Axis
car_counts <- mpg %>%
group_by(drv) %>%
summarize(total = n())
total_cars <- sum(car_counts$total)
ggplot(car_counts,
aes(x = drv, y = total,
color = drv)) +
geom_point() +
scale_y_continuous(
sec.axis = sec_axis(
trans = ~ . / total_cars,
labels = scales::percent,
name = "%")
) +
guides(fill = FALSE)+
theme_minimal()+
NULL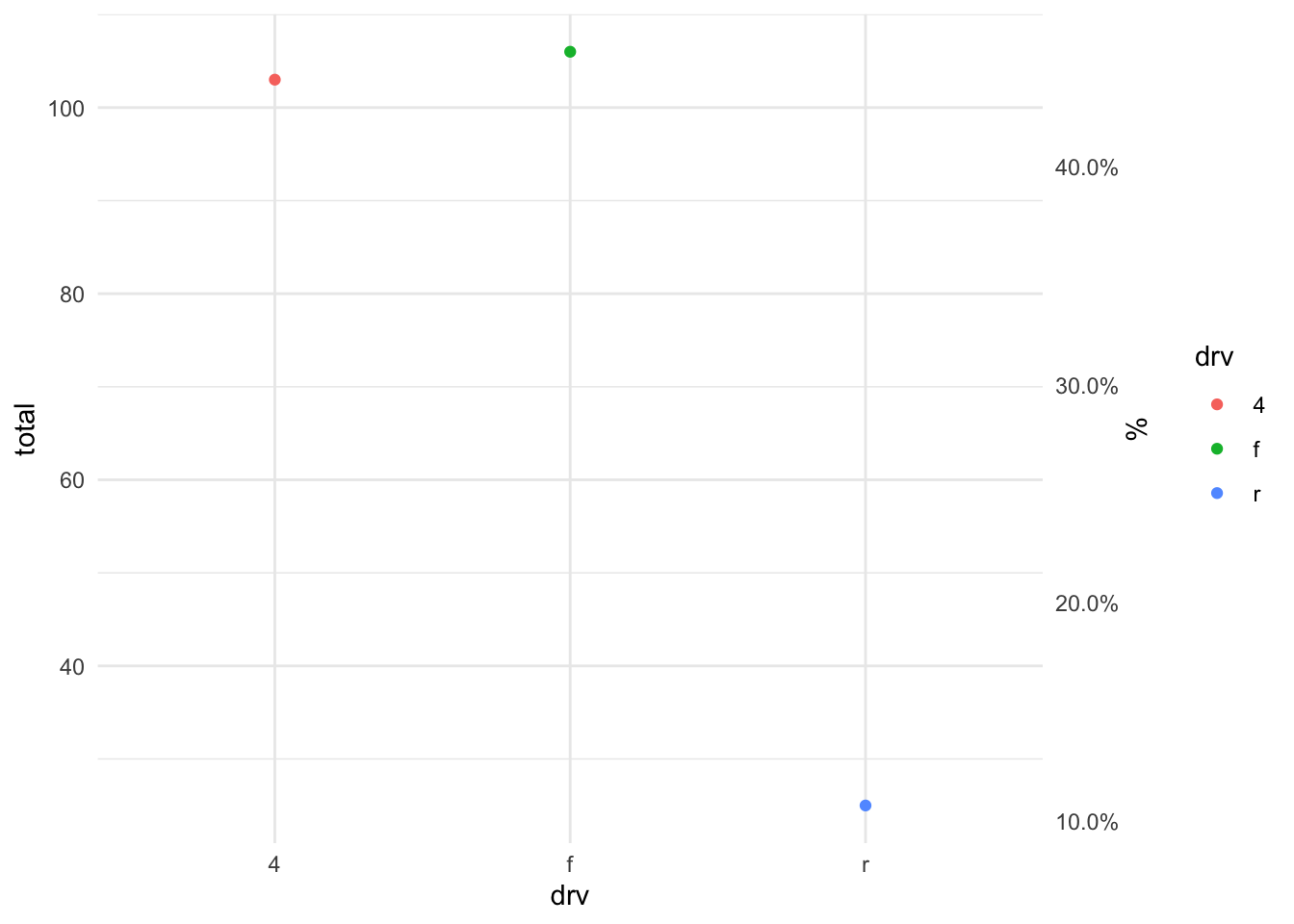
Time Series - Flights Number
library(nycflights13)
# just filter to have first 2 months
flights %>%
mutate(day=as.Date(time_hour)) %>%
filter(day < "2013-03-01") %>%
count(day,origin) %>%
ggplot(aes(x=day, y=n, color=origin)) +
geom_line(aes(group=origin)) +
geom_point() +
theme_bw()+
theme(legend.position="bottom")+
labs(subtitle="Number of Flights per Airports")+
NULL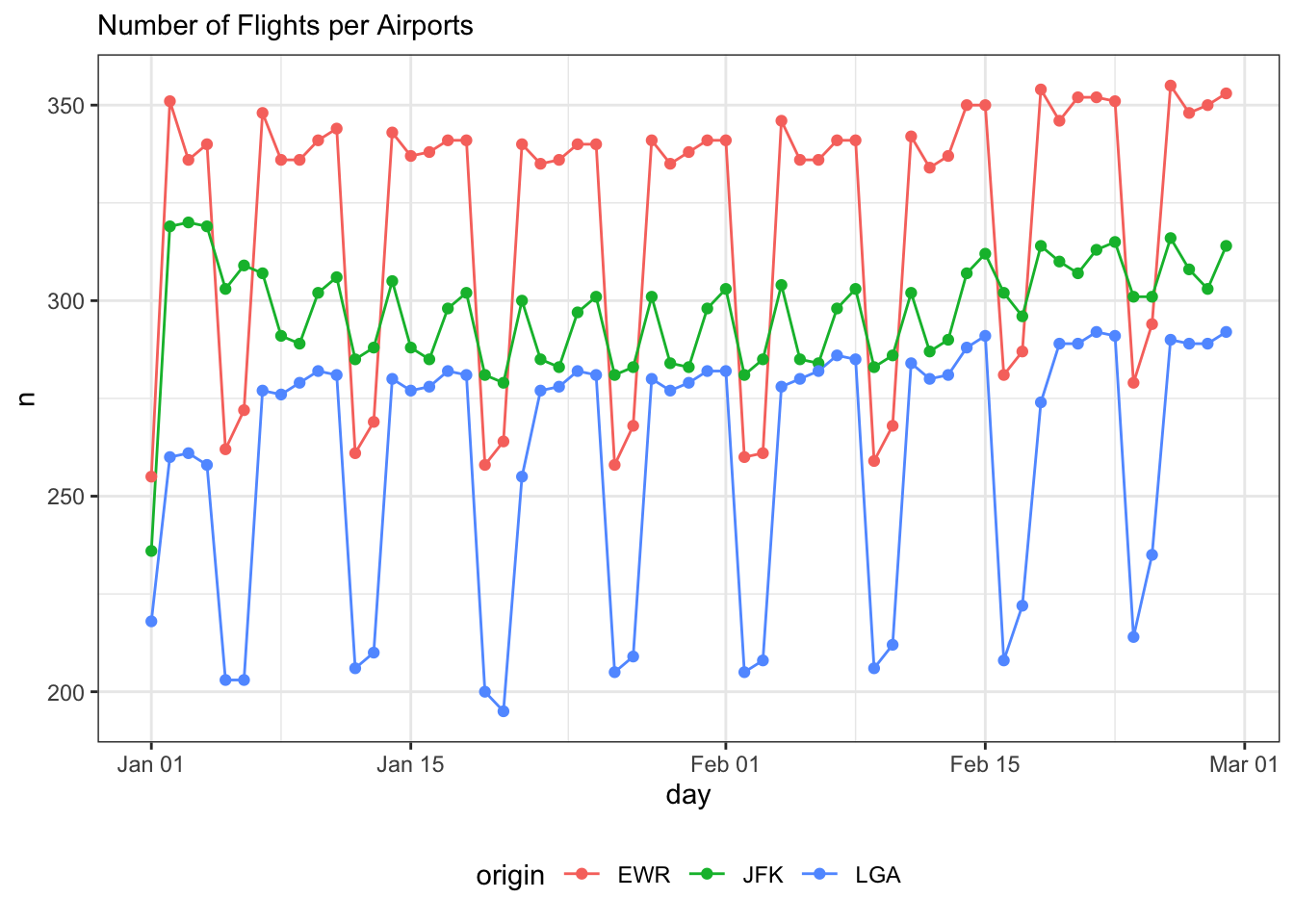
To highlight one among all others:
library(gghighlight)
africa <- gapminder %>%
filter(continent == "Africa") %>%
# sort by life expectancy in 1952, the first year in the dataframe
arrange(year, desc(lifeExp)) %>%
mutate(country = fct_inorder(factor(country)))
# find the African countries that had better life expectancy in 1992 compared to 2007
life_expectancy_dropped <- africa %>%
# pivot wider to be able to compare life expectancy in 1992 to 2007,
pivot_wider(country, names_from = year, values_from = lifeExp) %>%
# create new variables, `le_dropped`, that is `TRUE` if life expectancy was higher in 1992
# and `le_delta` which is simply the difference between life expectancy in 2007 and 1992
mutate(
le_delta = `2007` - `1992`,
le_dropped = `1992` > `2007`) %>%
select(-c(2:13))
# join `le_dropped` to each observation for each country
africa <- left_join(africa, life_expectancy_dropped, by = "country")
le_line_plot <- africa %>%
ggplot(aes(year, lifeExp, color = country, group = country)) +
geom_line(size = 1.2, alpha = 0.9, color = "#E58C23") +
theme_minimal(base_size = 14) +
# let us add a vertical line at 1992, when we start our comparison
geom_vline(xintercept = 1992, alpha = 0.1)+
theme(
# remove legend using the legend.position argument in theme()
legend.position = "none",
panel.grid.major.x = element_blank(),
panel.grid.minor = element_blank()
) +
labs(
title = "Life expectancy reduction, 1992 to 2007",
y = "life expectancy",
caption = "sorted by life expectancy in 1952") +
scale_x_continuous(breaks = seq(1952, 2007, 10))+
NULL
# Using `gghighlight()` we add add direct labels to the plot.
# The first argument defines which lines to highlight using `le_dropped`.
# Also add the arguments `use_group_by = FALSE` and `unhighlighted_colour = "grey80"`.
# If we facet_wrap() we do not need direct labels, so set `use_direct_label = FALSE`
le_line_plot +
gghighlight(
le_dropped,
use_group_by = FALSE,
use_direct_label = FALSE,
unhighlighted_colour = "grey80"
) +
facet_wrap(~country)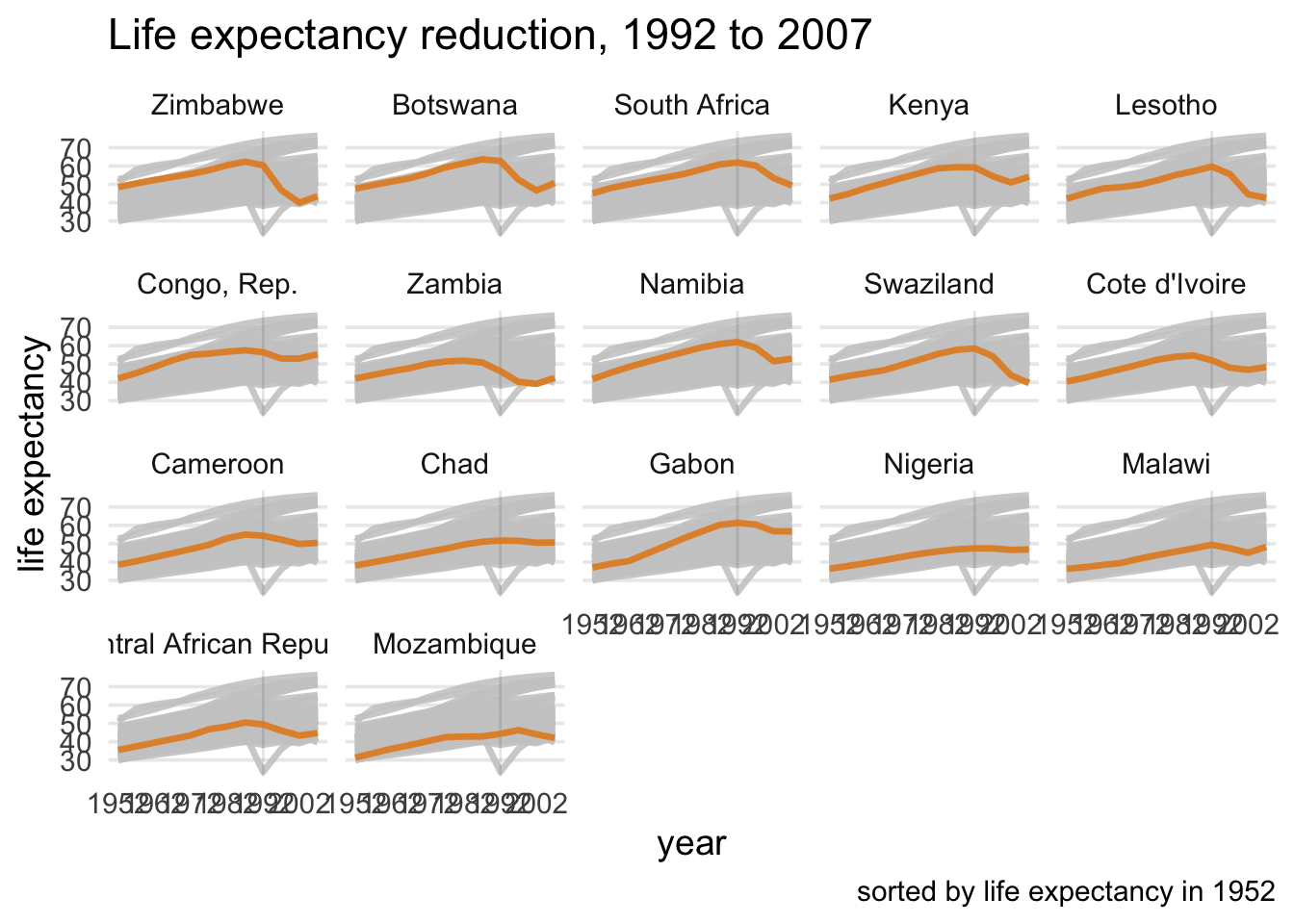
Confidence Interval
l3 <- c("compact","subcompact","midsize",
"2seater","minivan","suv","pickup")
# avg highway mpg with boostrapped 95% CI
mpg %>%
mutate(class = factor(class, levels = l3)) %>%
ggplot(aes(x = class, y = hwy, color = class))+
stat_summary(fun.y = mean, geom = "point") +
stat_summary(fun.data = mean_cl_boot,
geom = "errorbar") +
theme_bw() +
coord_flip() +
theme(legend.position = "none") +
labs(x = " ", y = "Highway MPG with 95% CI")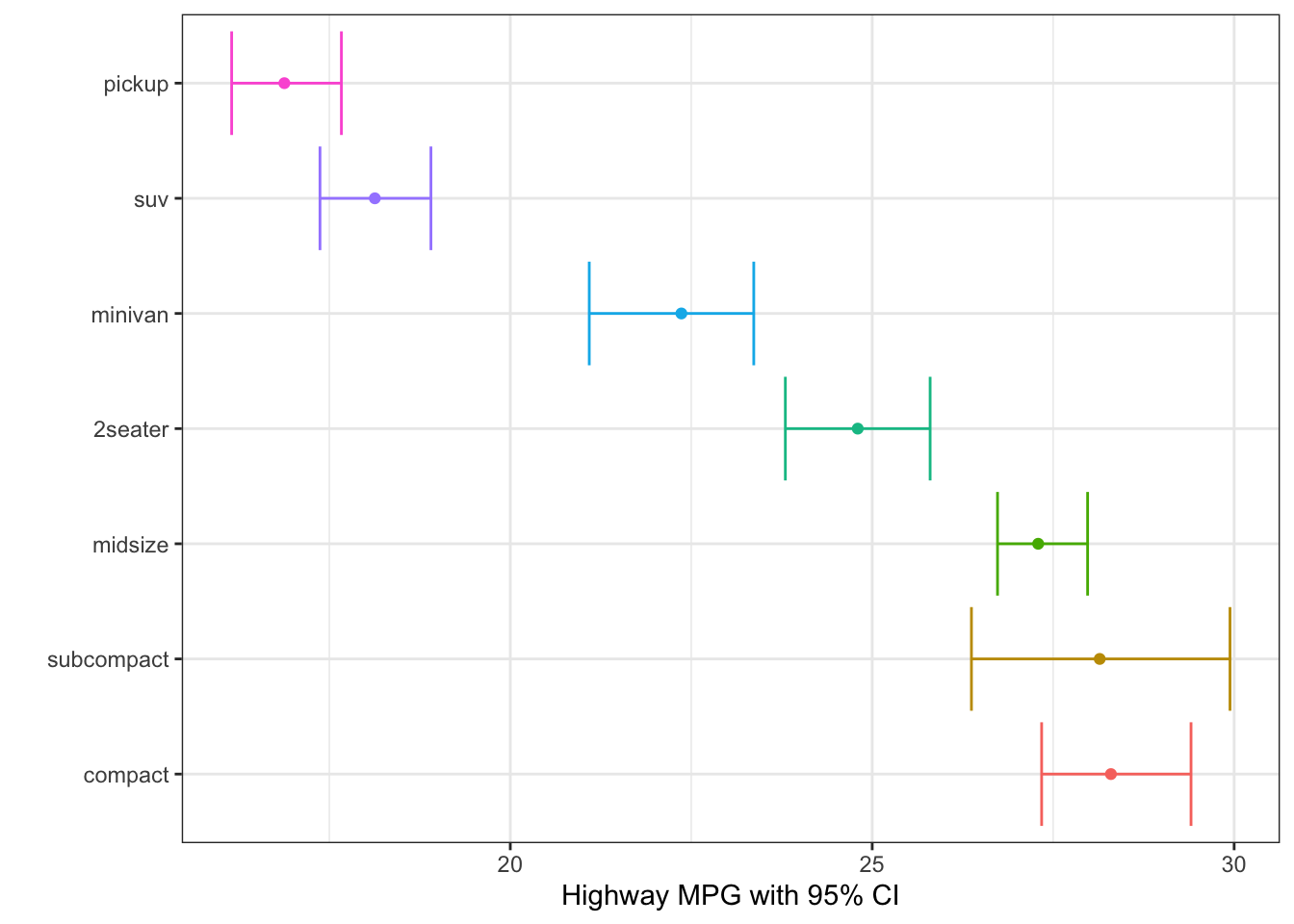
Jitter - to randomly spread the points
# geom_jitter
origin <- tibble(
x = rep(0, times = 10),
y = rep(0, times = 10)
)
#geom_point will only plot one point, as for all points x=0, y=0
ggplot(origin, aes(x=x, y=y))+
geom_point(size=5, colour="orange")
# geom_jitter will randomly spread the points around (0.0)
ggplot(origin, aes(x=x, y=y))+
geom_point(size=5, colour="orange")+
geom_jitter()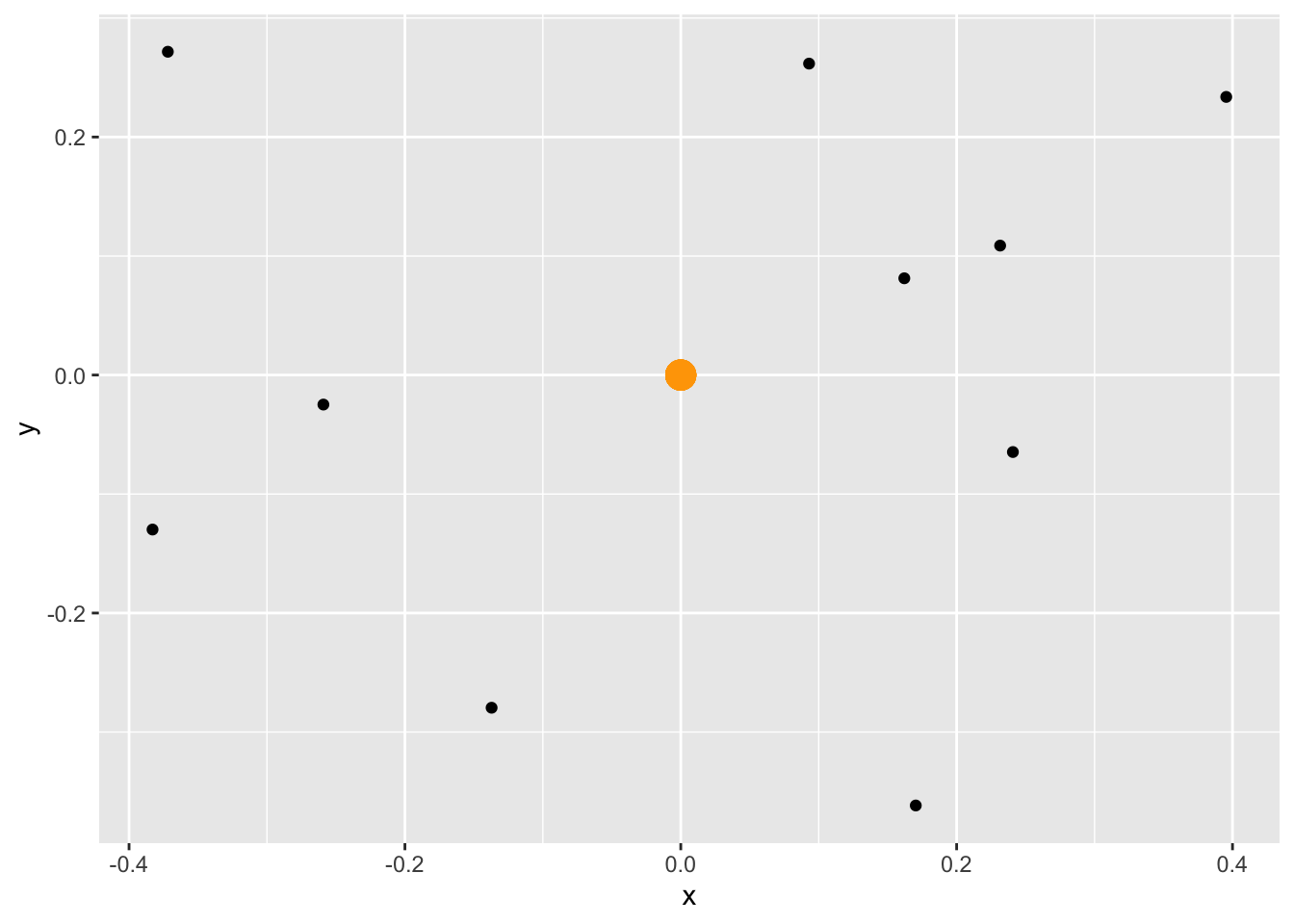
Maps
sf basics
library(sf)
library(usethis)
library(tidyverse)
library(rnaturalearth)
library(patchwork)
world <- ne_countries(scale = "medium", returnclass = "sf") %>%
filter(name != "Antarctica")
# what happens if we change the crs/projection
base_map <- ggplot() +
geom_sf(data = world,
aes(
geometry = geometry, #use Natural Earth World boundaries
fill = region_un #fill colour = country’s region
),
colour = "grey90", # borders between regions
show.legend = FALSE # no legend) +
)+
theme_minimal()+
NULL
base_map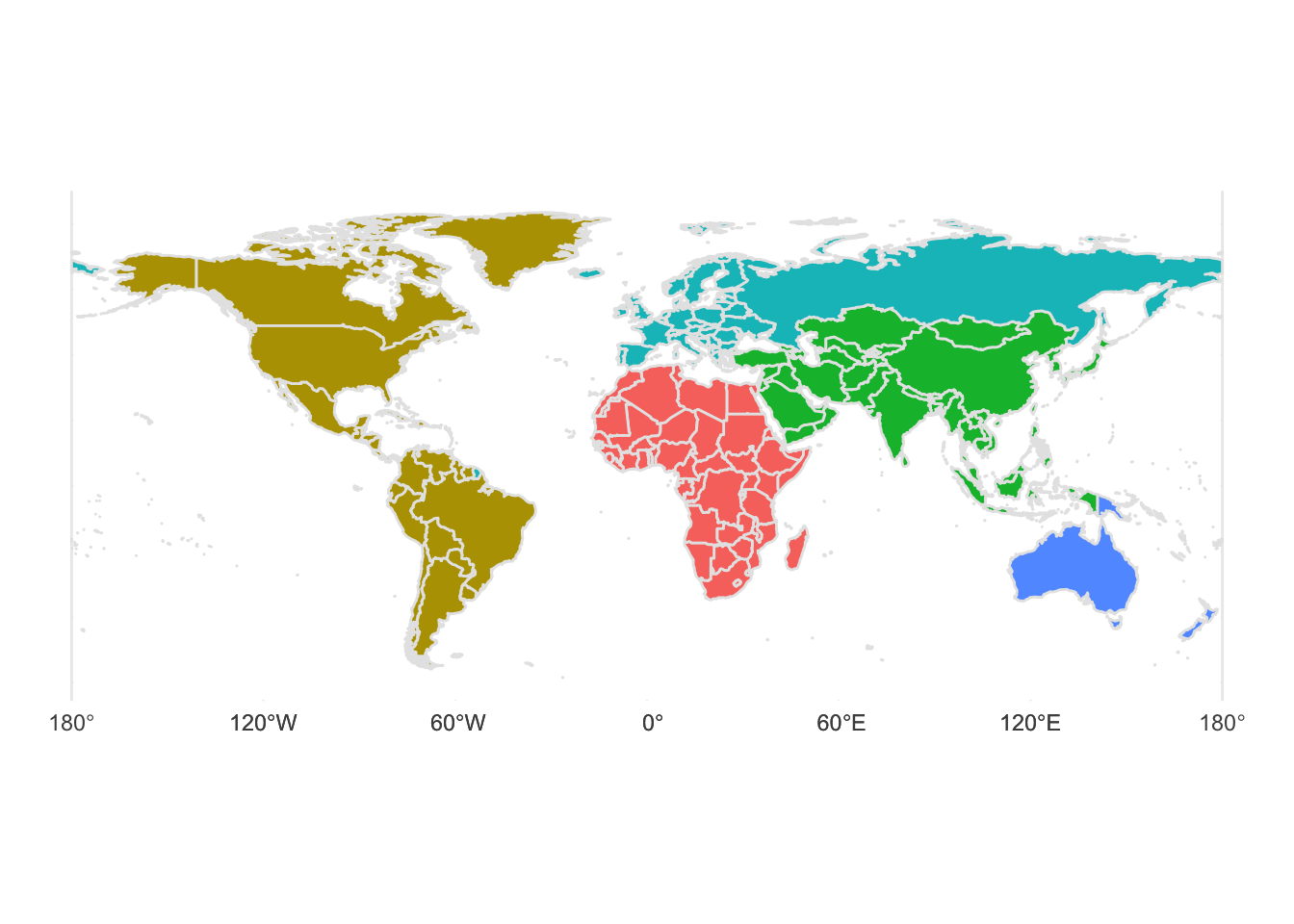
# Longitude/latitude
map_lat_lon <- base_map +
coord_sf(crs = "+proj=longlat +ellps=WGS84 +datum=WGS84 +no_def") +
labs(title = "Longitude-latitude",
subtitle = 'crs = "+proj=longlat +ellps=WGS84"')
# Robinson
map_robinson <- base_map +
coord_sf(crs = "+proj=robin") +
labs(title = "Robinson",
subtitle = 'crs = "+proj=robin"')
# Mercator (ew)
map_mercator <- base_map +
coord_sf(crs = "+proj=merc") +
labs(title = "Mercator",
subtitle = 'crs = "+proj=merc"')
# Azimuthal Equidistant
map_azimuthal_equidistant <- base_map +
coord_sf(crs = "+proj=aeqd") + # Gall Peters / Equidistant cylindrical
labs(title = "Azimuthal Equidistant",
subtitle = "crs = +proj=aeqd")
#use patchwork to arrange 4 maps in one page
(map_lat_lon / map_mercator) | ( map_robinson / map_azimuthal_equidistant)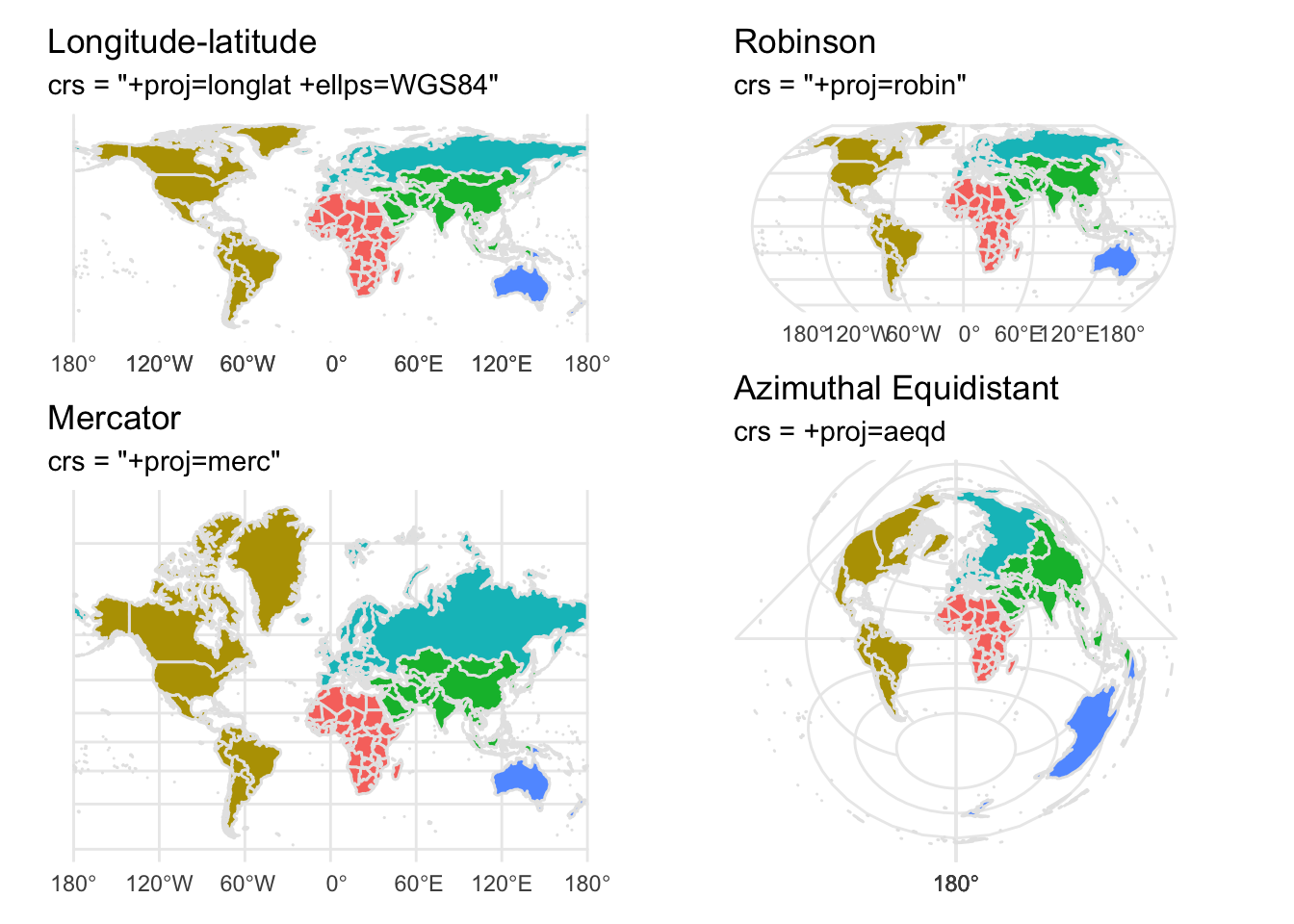
with Heatmap
# remotes::install_github("kjhealy/nycdogs")
library(nycdogs)
library(sf) # for geospatial visualisation
# load the data
data("nyc_license")
data("nyc_zips")
# what is the prevalent NYC breed ?
nyc_license %>%
count(breed_rc, sort=TRUE) ## # A tibble: 301 × 2
## breed_rc n
## <chr> <int>
## 1 Unknown 16797
## 2 Yorkshire Terrier 7786
## 3 Shih Tzu 7154
## 4 Labrador (or Crossbreed) 6986
## 5 Pit Bull (or Mix) 6751
## 6 Chihuahua 5785
## 7 Maltese 4308
## 8 Pomeranian 2197
## 9 Beagle 2111
## 10 Jack Russell Terrier 2041
## # … with 291 more rows# what is the prevalent NYC dog name by animal gender?
# Bella for girls, Max for boys!
nyc_license %>%
count(animal_name, animal_gender, sort=TRUE) ## # A tibble: 19,294 × 3
## animal_name animal_gender n
## <chr> <chr> <int>
## 1 Unknown M 1401
## 2 Bella F 1365
## 3 Max M 1272
## 4 Name Not Provided M 1111
## 5 Unknown F 1093
## 6 Lola F 875
## 7 Charlie M 871
## 8 Rocky M 869
## 9 Lucy F 767
## 10 Buddy M 743
## # … with 19,284 more rows# Maps for two Breeds
top_breeds <- nyc_license %>%
group_by(zip_code, breed_rc) %>%
tally() %>%
filter(n>10) %>%
mutate(freq = n / sum(n),
pct = round(freq*100, 2)) %>%
filter(breed_rc == "Yorkshire Terrier" | breed_rc == "Labrador (or Crossbreed)")
top_breeds_map <- left_join(nyc_zips, top_breeds) %>%
na.omit()
top_breeds_map %>% ggplot(mapping = aes(fill = pct)) +
geom_sf(color = "gray80", size = 0.1) +
scale_fill_viridis_b(option = "A", direction= -1) +
labs(fill = "Percent of Licensed Dogs") +
facet_wrap(~breed_rc)+
theme_void() +
guides(fill = guide_legend(title.position = "top",
label.position = "bottom")) 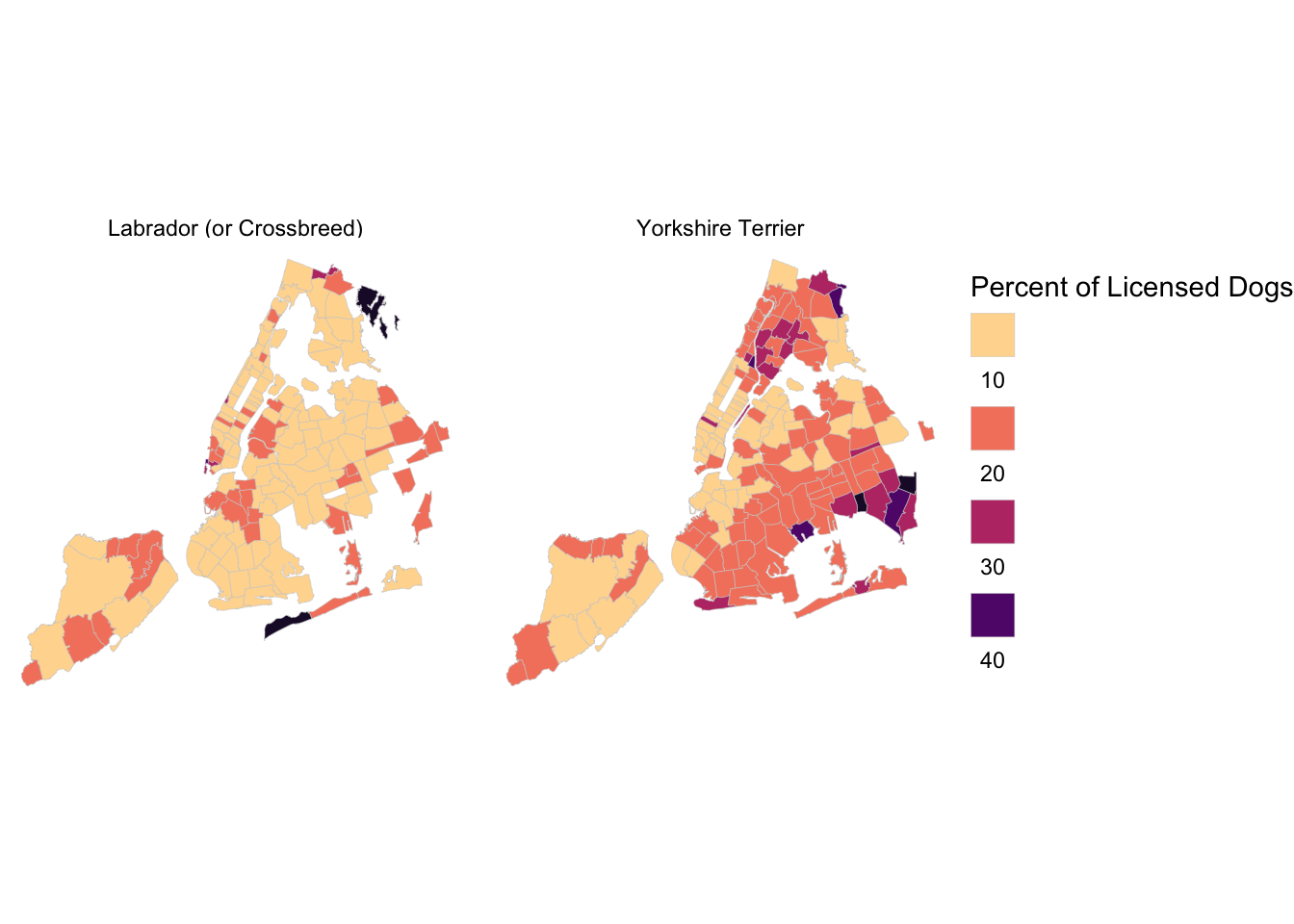
with Arrows
library(tidyverse)
library(sf)
library(rnaturalearth)
library(eurostat)
library(lubridate)
library(opencage)
library(hrbrthemes)
library(gridExtra)# Air passenger transport between the main airports of the United Kingdom
# and their main partner airports (routes data)
# https://ec.europa.eu/eurostat/web/products-datasets/-/avia_par_uk
flights_UK <- get_eurostat(id="avia_par_uk",
select_time = "M") #get monthly data
# filter passengers only
flights <- flights_UK %>%
dplyr::filter(unit == "PAS") %>%
mutate(
origin = str_sub(airp_pr, 1, 7),
destination = str_sub(airp_pr, -7),
from = str_sub(origin, 4,7),
to = str_sub(destination, 4,7),
year = year(time),
month_name = month(time, label = TRUE)
)
flights %>%
filter(year>=2018) %>%
group_by(origin,year) %>%
summarise(totalpassengers = sum(values)) %>%
arrange(desc(totalpassengers))## # A tibble: 95 × 3
## # Groups: origin [34]
## origin year totalpassengers
## <chr> <dbl> <dbl>
## 1 UK_EGLL 2019 315616656
## 2 UK_EGLL 2018 314171004
## 3 UK_EGKK 2019 174720542
## 4 UK_EGKK 2018 173091550
## 5 UK_EGCC 2019 102403412
## 6 UK_EGSS 2019 101440212
## 7 UK_EGSS 2018 100518160
## 8 UK_EGCC 2018 98376330
## 9 UK_EGLL 2020 64403150
## 10 UK_EGGW 2019 62841150
## # … with 85 more rowslondon_airports <- c("UK_EGLL","UK_EGKK","UK_EGSS","UK_EGGW")
london_flights <- flights %>%
dplyr::filter(origin %in% london_airports) %>%
distinct()
# count number of flights from origins to destinations
origins <- london_flights %>%
count(from, sort=TRUE) %>%
mutate(prop = n/sum(n))
destinations <- london_flights %>%
count(to, sort=TRUE) %>%
mutate(prop = n/sum(n))
# geocode origin/destination airports
origins <- origins %>%
mutate(
origin_geo = purrr::map(from, opencage_forward, limit=1)
) %>%
unnest_wider(origin_geo) %>%
unnest(results) %>%
rename(from_y = geometry.lat,
from_x = geometry.lng)
destinations <- destinations %>%
mutate(
destination_geo = purrr::map(to, opencage_forward, limit=1)
) %>%
unnest_wider(destination_geo) %>%
unnest(results) %>%
rename(to_y = geometry.lat,
to_x = geometry.lng)
#geocode airports
londonflights2018 <- london_flights %>%
filter(year==2018) %>%
group_by(year, destination, origin, from, to) %>%
summarise(totalpassengers = sum(values)) %>%
arrange(desc(totalpassengers)) %>%
#geocode origin (from_x, from_y)
mutate(
origin_geo = purrr::map(from, opencage_forward, limit=1)
) %>%
unnest_wider(origin_geo) %>%
unnest(results) %>%
rename(from_y = geometry.lat,
from_x = geometry.lng) %>%
select(destination, from, to, totalpassengers, from_x, from_y) %>%
#geocode destination (to_x, to_y)
mutate(
destination_geo = purrr::map(to, opencage_forward, limit=1)
) %>%
unnest_wider(destination_geo) %>%
unnest(results) %>%
rename(to_y = geometry.lat,
to_x = geometry.lng) %>%
select(destination, from, to, totalpassengers, from_x, from_y, to_x, to_y)
# for displaying a data table as annotation next to the map
table <- as_tibble(londonflights2018 %>% group_by(destination) %>% head(20)) %>%
select(Origin = origin, Destination = destination, Passengers = totalpassengers) %>%
mutate(Passengers = glue::glue("{scales::comma(Passengers)}")) %>%
tableGrob(
rows = NULL,
theme = ttheme_default(
core = list(
fg_params = list(
fontfamily = "Lato",
hjust = c(rep(0, 20), rep(1, 20)),
x = c(rep(0.1, 20), rep(0.9, 20))
)
)
)
)
ggplot() +
geom_sf(data = world, size = 0.125) +
geom_curve(
data = londonflights2018 %>% head(20),
aes(x = from_x, y = from_y, xend = to_x, yend = to_y,
size= totalpassengers, colour = totalpassengers),
curvature = 0.2, arrow = arrow(length = unit(3, "pt"), type = "closed"),
)+
theme_void()+
annotation_custom(table, xmin=-165, xmax=-160, ymin=-120, ymax=90) +
labs(
x = NULL, y = NULL,
title = "Top 20 destinations for London flights in 2018",
subtitle = "Data source: Eurostat , id=avia_par_uk"
) +
theme(
text=element_text(size=16, family="Lato"),
plot.title = element_text(),
plot.title.position = "plot",
axis.text.x = element_blank(),
axis.title.y = element_blank(),
legend.position = "none"
) +
NULL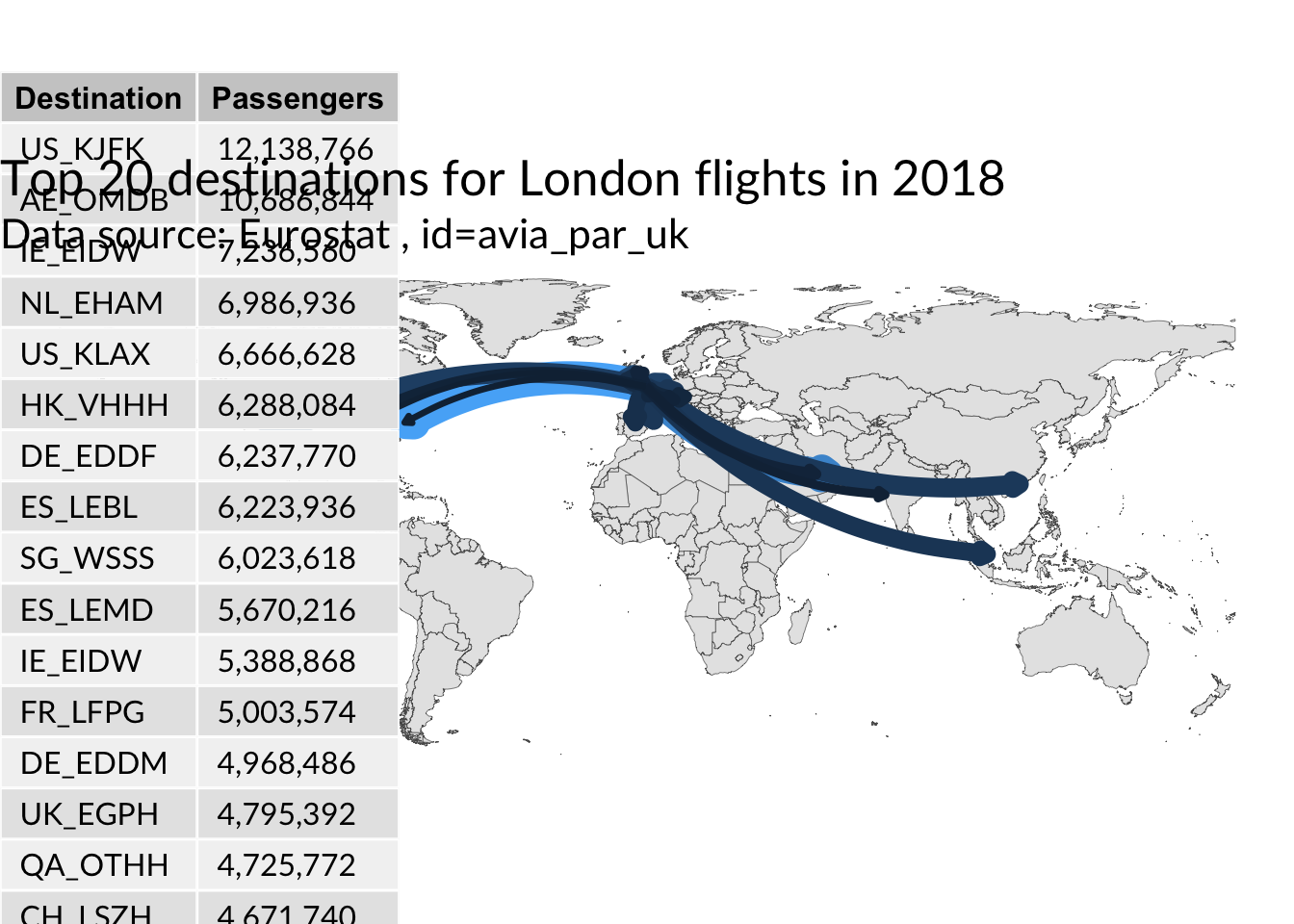
London Wards Map
mapview interactive map
library(tidyverse)
library(vroom)
library(sf)
library(rnaturalearth)
library(rnaturalearthdata)
library(rgdal)
library(rgeos)
library(patchwork)
library(mapview)
library(tmap)
library(viridis)url <- "https://covid.ourworldindata.org/data/owid-covid-data.csv"
covid_data <- vroom(url) %>%
mutate(perc = round(100 * people_fully_vaccinated / population, 2))
map <- ne_countries(scale = "medium", returnclass = "sf") %>%
dplyr::select(name, iso_a3, geometry) %>%
filter(!name %in% c("Greenland", "Antarctica"))
df <- map %>%
rename(iso_code=iso_a3) %>%
left_join(
covid_data %>%
select(location, iso_code, date, people_fully_vaccinated, perc, continent) %>%
group_by(location) %>%
slice_max(order_by = perc, n=1) %>%
ungroup(),
by = "iso_code") %>%
drop_na(date)
# use viridis plasma colour scale
df %>%
mapview::mapview(zcol = "perc",
at = seq(0, max(df$perc, na.rm = TRUE), 10),
legend = TRUE,
col.regions = plasma(n = 8),
layer.name = "Fully Vaccinated %")# Use tmap package
df %>%
filter(continent == "Asia") %>%
tm_shape() +
tmap_options(check.and.fix = TRUE)+
tm_polygons(col = "perc",
n = 5,
title = "% vaccinated",
palette = "Blues") 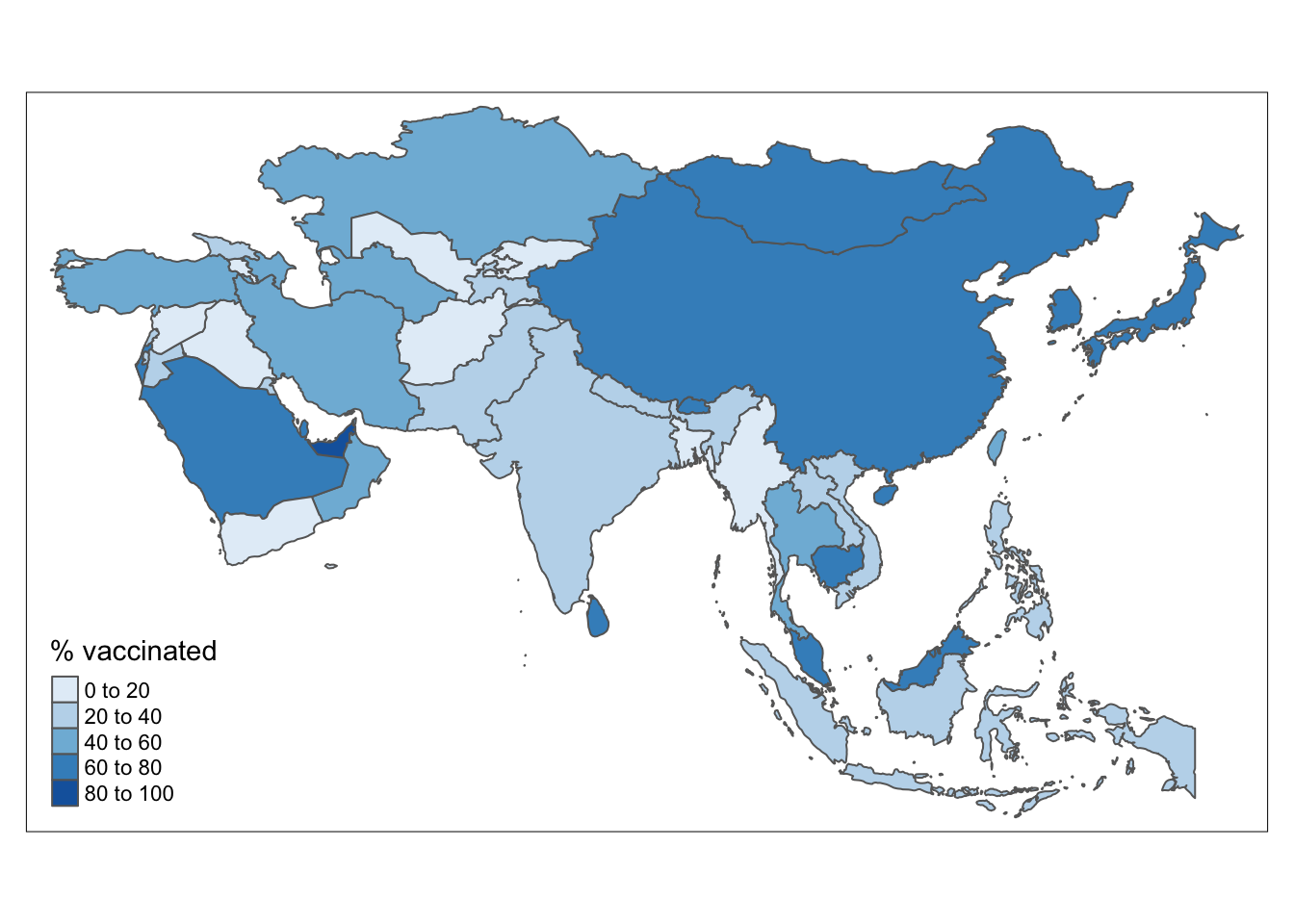
tmap_mode("view") # for interactive maps
df %>%
filter(continent == "Asia") %>%
tm_shape() +
tm_polygons(col = "perc",
n = 5,
title = "% vaccinated",
palette = "Blues") tmap_mode("plot") # back to static maps for tmap
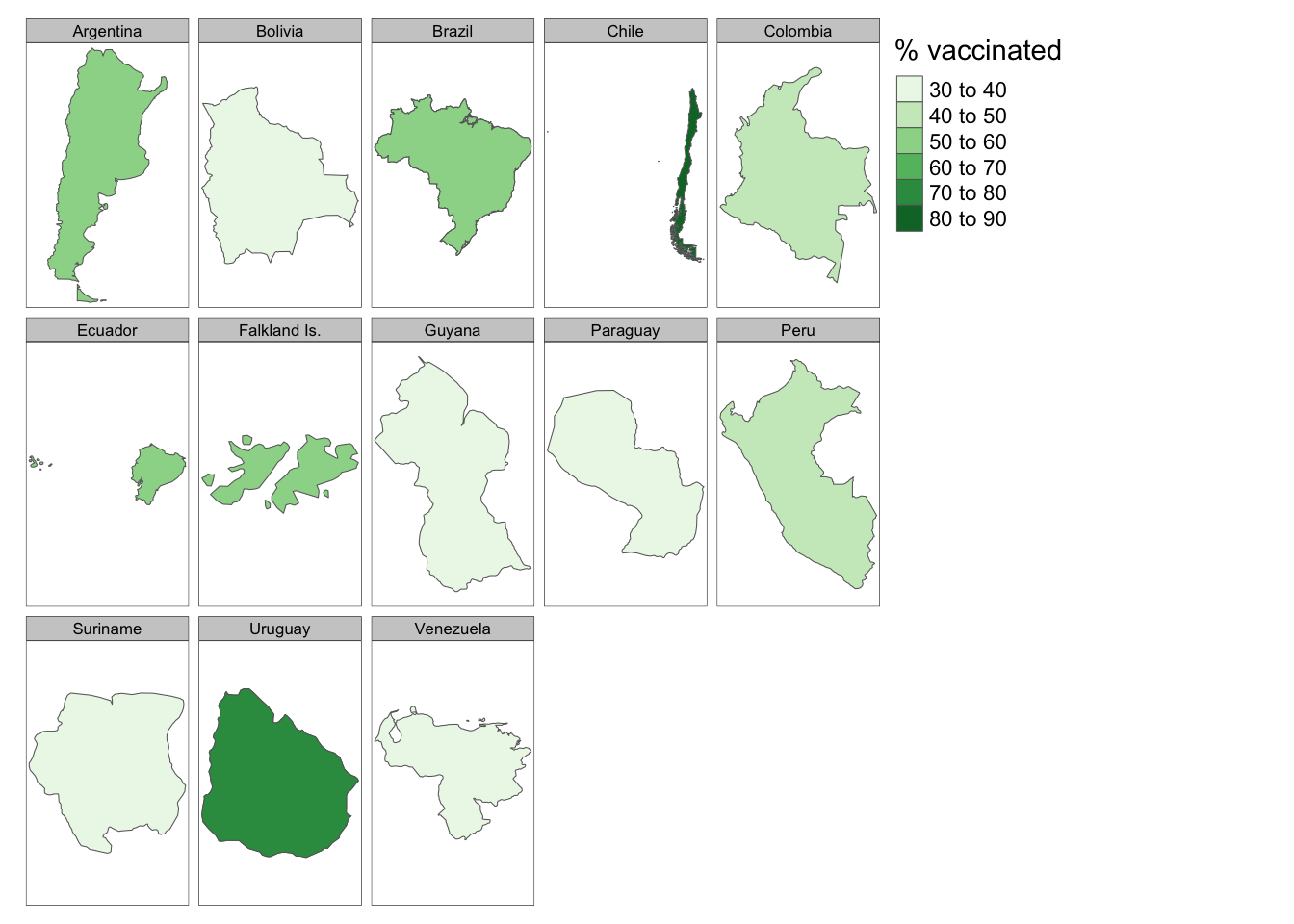
df %>%
filter(continent == "South America") %>%
tm_shape() +
tm_polygons(col = "perc",
n = 5,
title = "% vaccinated",
palette = "Greens") +
tmap_options(limits = c(facets.view = 13)) +
tm_facets(by = "name")